Figure 3.
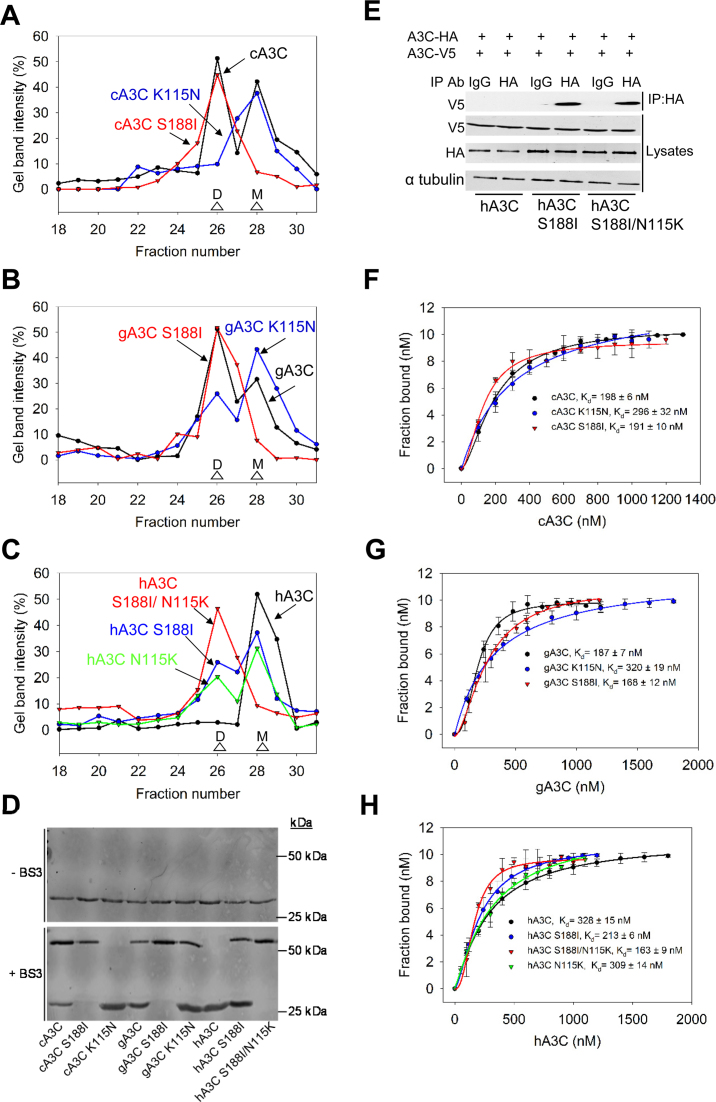
A3C dimerization is mediated through α-helix 6 or β-strand 4. (A–C) SEC profile for 10 μg of (A) cA3C, cA3C K115N and cA3C S188I; (B) gA3C, gA3C K115N and gA3C S188I and (C) hA3C, hA3C S188I, hA3C N115K and hA3C S188I/N115K from a 10 ml Superdex 200 column was used to calculate the oligomerization state of the enzyme from a standard calibration curve. An M denotes a monomer fraction and a D denotes a dimer fraction. (A) cA3C formed monomers and dimers (apparent molecular weights 19 kDa and 45 kDa, respectively), cA3C S188I formed a stable dimer (apparent molecular weight 45 kDa), and cA3C K115N formed monomers (apparent molecular weight 19 kDa). (B) The gA3C SEC profiles were similar to cA3C, except for gA3C K115N that was mainly monomers (apparent molecular weight 19 kDa), but also retained a small proportion of dimers (apparent molecular weight 45 kDa). (C) hA3C formed monomers in solution (apparent molecular weight 19 kDa), hA3C S188I and hA3C N115K were an equilibrium of monomers and dimers (apparent molecular weights 19 kDa and 45 kDa, respectively) and hA3C S188I/N115K was a stable dimer (apparent molecular weight 45 kDa). The chromatograms were constructed by analyzing the integrated gel-band intensities of each protein in each fraction after resolution by SDS-PAGE (Supplementary Figure S3). (D) A3C enzymes were incubated in the absence or presence of 20 μM BS3 crosslinker and subsequently visualized with SDS-PAGE and immunoblotting. Monomeric A3C enzymes remained as monomers in the presence of crosslinker (cA3C K115N, hA3C). A3C enzymes that were able to form dimers according to SEC, were also present as monomers/dimers (cA3C, gA3C, gA3C K115N, hA3C S188I) or as dimers (cA3C S188I, gA3C S188I, hA3C S188I/N115K) in the presence of the crosslinker. Molecular weight standards are indicated. (E) Coimmunoprecipitation of A3C-V5 with A3C-HA. The A3C-HA and A3C-V5 were transfected in combination and the immunoprecipitation of cell lysates used either anti-HA antibody or Rabbit IgG (mock) and was immunoblotted with antibodies against α-tubulin, HA and V5. Cell lysates show the expression of α-tubulin, HA and V5. (F–H) The apparent Kd of A3C enzymes from a 118 nt ssDNA was analyzed by steady-state rotational anisotropy for (E) cA3C, cA3C S188I and cA3C K115N; (F) gA3C, gA3C S188I, and gA3C K115N and (G) hA3C, hA3C S188I, hA3C N115K and hA3C S188I/N115K. Apparent Kd values are shown in the figure. Hill coefficients for cooperative binding curves are (E) cA3C, 1.6; cA3C S188I, 1.7; (F) gA3C, 1.8; gA3C S188I, 2.1; (G) hA3C S188I, 1.6; hA3C N115K, 1.5; hA3C S188I/N115K, 1.9. Error bars represent the S.D. from three independent experiments.