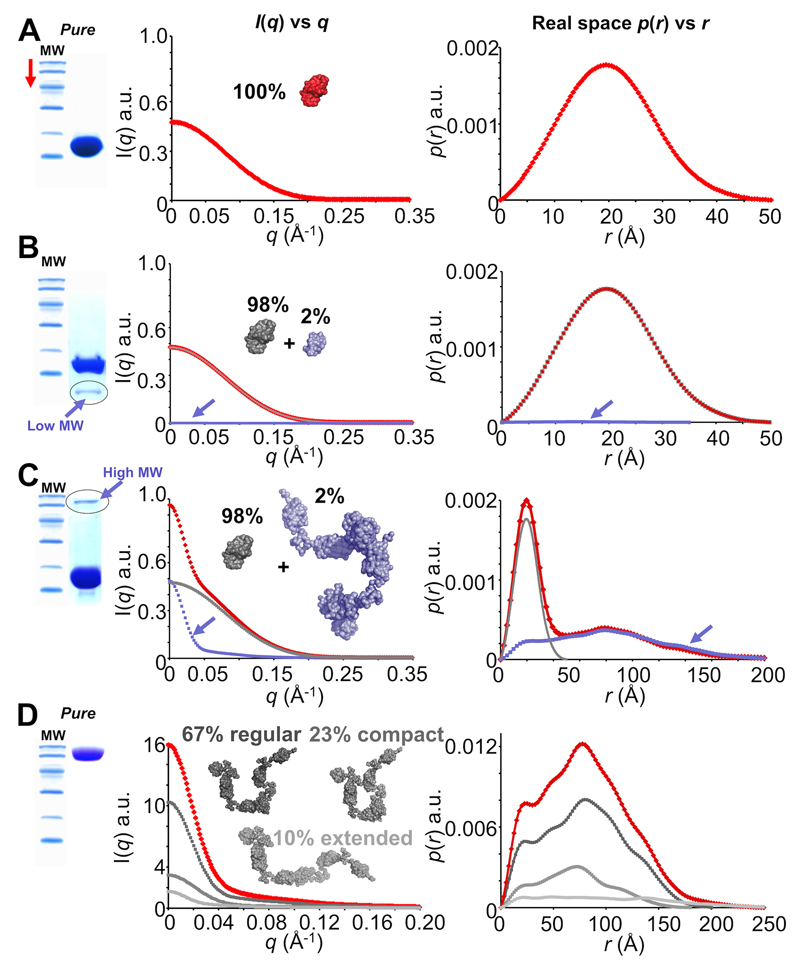
Figure 6. Sample purity and contaminants (simulated SAXS data and simulated SDS-PAGE).
A. The ideal outcome when purifying a sample. Scattering from each individual within a population of pure monodisperse 14 kDa protein sum to produce a total scattering profile (red) from which p(r) vs r can be modelled that represents the real-space atom-pair distance distribution within a single particle. B. A less-ideal situation. If contaminants are present, the total scattering (red) will be comprised of the sum of the scattering from each different species in proportion to their volume squared and concentration. Here, a low molecular weight (MW) contaminant (~5 kDa, 2% of the sample, blue) is present in the 14 kDa protein sample (grey). However, the total contribution to the scattering made by the low MW contaminant is small and does not significantly affect I(q) vs q or p(r) vs r. C. Something to avoid. High MW contaminants have disastrous consequences on I(q) vs q (red). The scattering contributions made by trace ~100 kDa protein (blue) doubles I(0) even though the target 14 kDa protein (grey) is 98% pure. The effect on p(r) vs r is significant as it is the sum-weighted contribution made by the 14 kDa protein plus the 100 kDa contaminant. D. A special case: flexibility. A 100 kDa protein is both pure and monomeric. However, the protein is flexible and is comprised of three main populations so that the total P(q) determined from the scattering (red) is the sum of P(q) from each population (shades of grey). For example, although the extended state comprises only 10 % of the total population, the maximum dimension of the measured p(r) vs r (red) will equate to the Dmax of the most extended state (light grey). Note before: the SDS-PAGE gels and scattering profiles used for this figure are for illustrative purposes only and do not represent real data.