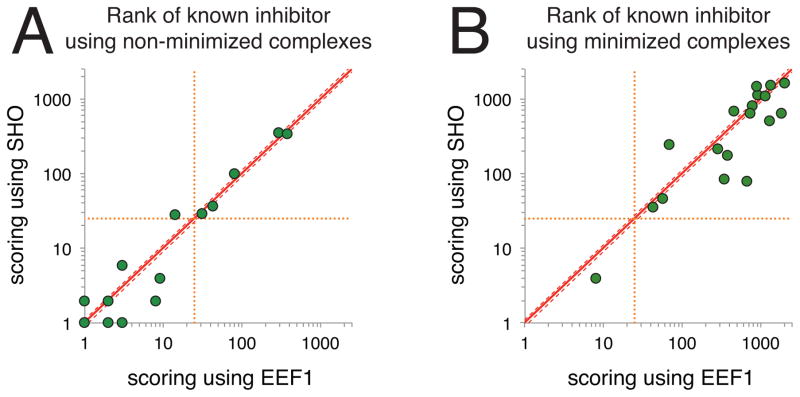
Figure 4. Applying SHO to identify small-molecule inhibitors of protein–protein interactions.
For each of 18 target proteins, 2500 “decoy” compounds were docked to the protein interaction site; the energies of these decoys were compared to that of a known “active” compound for each complex (the active compound differed for each target protein, the complete collection is shown in Figure S1). For each of these complexes and their corresponding native complex, the interaction energy was determined using the Rosetta energy function with EEF1 or using the Rosetta energy function with SHO. For each target, we report the rank of the active compound (i.e., the native complex) relative to the decoys; each point on the plots represents a different protein target. Points below the diagonal represent targets for which SHO ranked the active compound ahead of more decoy compounds than EEF1. A) In the initial benchmark, the protein–ligand complexes were not pre-minimized. Excluding “ties” (cases in which the ranks for both methods were within 10% of one another), SHO outperforms EEF1 for 7 targets whereas EEF1 outperforms SHO for 5 targets. Only 6 points are visible for the former, because there are two targets for which the rank by SHO is 1 and the rank by EEF1 is 2. B) In the subsequent experiment, each of the protein–ligand complexes was pre-minimized using the standard Rosetta energy function. In this more challenging benchmark, SHO outperforms EEF1 for 11 targets whereas EEF1 outperforms SHO for 5 targets.