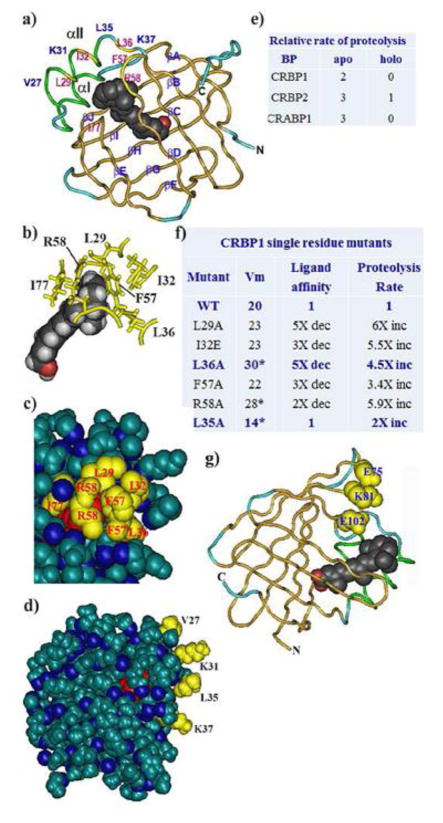
Fig. 7.
Structure-activity relationships of retinoid BP. a) Ribbon diagram of CRBP1 illustrating the entrance portal, key residues, orientation of retinol in the binding pocket, and the structure of the β-clam (Cowan et al., 1993; Franzoni et al., 2002; Lu et al., 2003; Careri et al., 2006; Mittag et al., 2006; Franzoni et al., 2010; 2016 Silvaroli et al., 2016). Red shows the hydroxyl group of retinol. Structure PDB ID 5HBS, NCBI. b) Partial structure focusing on interactions of entrance portal residues with each other and with the β-ionone ring. Red depicts the retinol hydroxyl group. c) Partial space-filling structure illustrating close contacts of residues in the entrance portal. Red illustrates the β-ionone ring. d) Total space-filling structure of CRBP1 showing sequestration of retinol from the cellular milieu, and residues that project from the αII helix out from the BP, which serve as putative “handles” that allow enzymes to tease retinol from the BP. Red indicates the β-ionone ring. e) Differences in proteolysis rates between apo- and holo-BP. BP were exposed to the endopeptidase Arg-C (clostripain). Degrees of proteolysis were analyzed by SDS-PAGE: scale: 0, no proteolysis; 1, low (<50%); 2, moderate (>50%); 3, complete. These data were obtained after 1 hour incubation, but holo-forms remained protease resistant up to 20 hours (Jamison et al., 1994). f) Effects of point mutations on ability of CRBP1 to deliver retinol to RDH, ligand affinity and rigidity (proteolysis rate) (Penzes and Napoli, 1999). g) Structure of CRABP1, which folds similarly to CRABP2, showing the carboxyl group (red) of atRA and the location of the three residues that enable RAR binding in CRABP2. Structure PDB ID 1CBR, NCBI.