Abstract
Algae are smaller organisms than land plants and offer clear advantages in research over terrestrial species in terms of rapid production, short generation time and varied commercial applications. Thus, studies investigating the practical development of effective algal production are important and will improve our understanding of both aquatic and terrestrial plants. In this study we estimated multiple physicochemical and secondary structural properties of protein sequences, the predicted presence of post-translational modification (PTM) sites, and subcellular localization using a total of 510,123 protein sequences from the proteomes of 31 algal and three plant species. Algal species were broadly selected from green and red algae, glaucophytes, oomycetes, diatoms and other microalgal groups. The results were deposited in the Algal Protein Annotation Suite database (Alga-PrAS; http://alga-pras.riken.jp/), which can be freely accessed online.
Keywords: Algae, Comparative analysis, Database, Gene function, Protein properties
Introduction
Algae are smaller organisms than land plants and offer clear advantages over terrestrial species for use in research in terms of rapid production, short generation time and varied commercial applications. Thus, algae are a very promising group of organisms for potential commercial applications, such as food and feed production, nutritional supplements, biofuel sources and environmental improvement through hydrogen production (Wijffels and Barbosa 2010, Draaisma et al. 2013, Torzillo et al. 2015). In the algal food and nutritional supplement sector, Chlorella vulgaris and Spirulina platensis have already been commercialized as health foods (Beheshtipour et al. 2013, Borowitzka 2013). However, while several studies in the biofuel sector have investigated selection, cultivation, extraction and purification of specific algal species and strains (Carvalho et al. 2006, Chisti 2007), a consensus has not yet been reached on costs and best practices in algal production (Passell et al. 2013, Medipally et al. 2015). Thus, studies investigating the development of practical and effective algal production techniques are important, and will improve our understanding of both aquatic and terrestrial plants, considering that algae are common ancestors of vascular plants (Reijnders et al. 2014, Bhattacharya et al. 2015).
The entire nuclear genome sequences of the red alga Cyanidioschyzon merolae (Matsuzaki et al. 2004) and the diatom Thalassiosira pseudonana (Armbrust et al. 2004) were determined. Subsequently, next-generation applications, including sequence assembly tools and gene prediction tools, have enabled the sequencing of algal species (Kim et al. 2014). As a result, over 30 whole algal genomes have been sequenced to date (Kim et al. 2014, Reijnders et al. 2014). These representative genomes, except for those of the two species mentioned above, include the green algae Ostreococcus tauri (Derelle et al. 2006) and Chlamydomonas reinhardtii (Merchant et al. 2007) of the Viridiplantae kingdom (including green plants), the red alga Galdieria sulphuraria (Schonknecht et al. 2013) and the glaucophyte Cyanophora paradoxa (Price et al. 2012). Additionally, genomes of the diatoms Phaeodactylum tricornutum (Chromista) (Bowler et al. 2008), Aureococcus anophagefferens (Pelagophyceae) (Gobler et al. 2011), Ectocarpus siliculosus (Phaeophyceae) (Cock et al. 2010), Emiliania huxleyi (Haptophyceae) (Read et al. 2013) and Guillardia theta (Cryptophyceae) (Curtis et al. 2012) are also included.
There is a considerable amount of information about land plants based on genomic, transcriptomic, proteomic and metabolomic analyses. The land plant Arabidopsis thaliana is currently one of the most commonly used experimental plants, as it has a small genome and a short life cycle. Information on Arabidopsis research was organized into The Arabidopsis Information Resource (TAIR) (Berardini et al. 2015). Similarly, Oryza sativa, also a well-studied species, is one of the most important crop plant models. Information regarding the genome and functional gene annotations in O. sativa is housed in the Michigan State University Rice Genome Annotation Project database (MSU Rice) (Ouyang et al. 2007) and the Rice Annotation Project database (RAP-DB) (Sakai et al. 2013). Furthermore, the genomic sequence information of various plant species has been updated in the JGI Genome Portal (Nordberg et al. 2014), Phytozome (Goodstein et al. 2012), GRAMENE (Youens-Clark et al. 2011) and PlantGDB (Dong et al. 2004). Moreover, in order to promote the development of functional annotation of genes in plants, several approaches and databases have been developed, accruing information on the transcriptome or metabolome in plants, as follows: transcription factor (TF) annotation at both family and gene levels (PlantTFDB) (Guo et al. 2008), TF integration of gene expression data for plants (ATTED-II) (Aoki et al. 2016b), integrative analysis for plant hormone accumulation and gene expression in rice (UniVIO) (Kudo et al. 2013), and utilization of transcriptomic and metabolic profiles among plant tissues (PRIMe Update) (Sakurai et al. 2013). These databases can be used to study gene function. Several large-scale experimental and computational approaches have also been adopted to enhance the study of functional annotation in plant proteomes (Kourmpetis et al. 2011, Akiyama et al. 2014, Clemente and Jamet 2015, Kurotani et al. 2015).
In algae, many general resources and culture collection databases exist, including: AlgaTerra (http://www.algaterra.org), AlgaeBase (http://www.algaebase.org) (Guiry et al. 2014), SAG (http://www.uni-goettingen.de/en/184982.html), NIES (http://mcc.nies.go.jp), and KU-MACC (http://www.research.kobe-u.ac.jp/rcis-ku-macc/E.index.html). Concomitantly, molecular-based biological approaches to algae have also been systematically recorded and made available through databases. These are: the database of genomic information of photosynthesis (Pico-PLAZA) (Vandepoele et al. 2013), the database of algal gene expression (ALCOdb) (Aoki et al. 2016a), the Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP) (Keeling et al. 2014), the database of Pleurochrysis transcripts (Pleurochrysome) (Yamamoto et al. 2016), the database of algal metabolic pathways (ALGAEpath) (Zheng et al. 2014) and the metabolome analyses of Cyanidioschyzon merolae (Sumiya et al. 2015). Although biological information on algae has been steadily increasing through research, it is still insufficient to comprehensively understand the functional annotations of algal genes.
Chlamydomonas reinhardtii is one of the best-studied green algae of recent years (May et al. 2009, Blaby et al. 2014, Aoki et al. 2016). According to the UniProt database (Bateman et al. 2015), as of July 2016 there were 14,716 records of C. reinhardtii. However, two-thirds of these records (9,860 records) are not informative annotations (e.g. ‘Predicted protein’, ‘Predicted protein -Fragment-’, and ‘Uncharacterized protein’) and only a subset of fewer than 50 annotations have experimentally validated functions (Reijnders et al. 2014). Therefore, comprehensive algal proteome information is far from satisfactory. Here we report the development of the Algal Protein Annotation Suite (Alga-PrAS) database, a user-friendly website with algal proteome information, specifically physicochemical, structural and functional annotations of algal proteome data.
Results and Discussion
Protein sequence sets
To provide unbiased proteome information, we prepared non-redundant protein sequence sets from whole-protein sequence sets of 34 species as follows. Sequences with fewer than 50 amino acids were omitted as these short sequences typically define peptides (Orlowski and Bujnicki 2008, Saghatelianr and Couso 2015). To avoid calculation failure of analytic tools, such as DIpro (Cheng et al. 2006), SSpro (Cheng et al. 2005) and DROP (Ebina et al. 2011), we removed sequences with more than 4,000 amino acids. Redundant sequences were removed by individually clustering protein sequences of each species. This was performed with the CD-HIT program (Fu et al. 2012) with default runtime options. Finally, 34 non-redundant protein sequence sets were independently obtained, totaling 510,123 sequences (Supplementary Table S1).
Annotation of algal proteomes by sequence similarity against public databases
Nonredundant algal protein sequences were aligned with BLASTP (Altschul et al. 1997, Altschul et al. 2005) against UniProtKB (Bateman et al. 2015). As a result, 46.2% of the algal protein sequences could achieve a hit with an e-value lower than 1e−10 (Table 1). The hit sequence percentages of 14 algae did not reach 50% (Supplementary Table S2). Approximately 60% of the algal proteins were annotated successfully, even when all assignment results to public databases were totaled. These results imply that functional genomic investigations are less efficient in algae than in land plants. Therefore, in addition to sequence similarity, the functional annotation of algal genomes should be enhanced by analytic approaches that employ structural and physicochemical properties, and post-translational modification (PTMs).
Table 1.
Percentages of sequences annotated by the KOG, Pfam, UniProtKB, GO and PDB databases
| Class | Percentage of annotated sequencesa (%) | |||||
|---|---|---|---|---|---|---|
| KOG | Pfam | UniProtKB | GO | PDB | Totalb (%) | |
| Land plants | 34.2 | 67.9 | 70.7 | 44.9 | 47.4 | 77.3 |
| Algae | 26.9 | 54.6 | 46.1 | 34.7 | 36.6 | 60.3 |
| Green algae | 31.8 | 60.7 | 55.9 | 38.9 | 41.7 | 67.3 |
| Red algae | 34.6 | 61.5 | 55.7 | 41.0 | 44.0 | 67.1 |
| Glaucophyceae | 14.0 | 31.4 | 25.7 | 19.5 | 19.8 | 37.0 |
| Oomycetes | 28.6 | 57.8 | 49.6 | 37.8 | 38.5 | 64.2 |
| Diatoms | 25.1 | 53.7 | 39.8 | 33.7 | 34.2 | 57.9 |
| Other microalgae | 22.8 | 50.8 | 39.1 | 31.2 | 33.4 | 56.0 |
| All species | 28.0 | 56.5 | 49.6 | 36.2 | 38.2 | 62.8 |
a Poor annotations such as ‘poorly characterized’ in KOG, ‘domain unknown function (DUF)’ in Pfam, and ‘Uncharacterized protein,’ ‘Putative uncharacterized,’ ‘Unnamed product’ and only ID in UniProtKB, were excluded from hits.
b Values were calculated by combining the results of KOG, Pfam, UniProtKB, GO and PDB.
Protein property information of Alga-PrAS
Compared with higher organisms, such as Homo sapiens (Imanishi et al. 2004), Mus musculus (McGarvey et al. 2015) and Arabidopsis (Berardini et al. 2015), available information and tools for the comprehensive annotation of algal proteomes are scarce. Therefore, it is important to provide information on algal protein function, specially that relating to protein properties. Physicochemical properties are useful to understand fundamental aspects of the structural stability, reactivity and solubility of proteins. Structural properties aid in identifying protein secondary structure and functional annotations against other existing protein sequences that are assigned to structural and functional domains or regions. In addition, PTM and subcellular localization aid in elucidating potential protein diversity, structure and function. We estimated 28 protein properties to improve the information on algal protein function with respect to various protein properties as stated above (Table 2). All information on the protein properties was integrated and housed in the Alga-PrAS database.
Table 2.
List of calculated protein properties in this study
| Classification of protein properties | Sub-classification of protein properties |
|---|---|
| Physicochemical properties | Protein length |
| Percentage of charged residues | |
| Percentage of nonpolar residues | |
| Percentage of acidic residues | |
| Percentage of basic residues | |
| Grand average value of hydropathicity index (GRAVY) | |
| Isoelectric point (pI) | |
| Probability of protein solubility | |
| Structural properties | Percentage of beta-pleated sheet secondary structure |
| Percentage of disordered residues | |
| Number of long disordered regions | |
| Existence of signal peptide cleavage site | |
| Number of transmembrane helices | |
| Number of S–S bonds | |
| Number of domain linkers | |
| Number of internal repeats | |
| Number of PEST regions | |
| Post-translational modifications (PTMs) and subcellular localization | Number of Ser, Thr and Tyr phosphorylation sites |
| Number of O-linked glycosylation sites | |
| Number of N-linked glycosylation sites | |
| Number of ubiquitination sites | |
| Protein subcellular localization sites |
Search interface of Alga-PrAS
We developed a proteome annotation database, Alga-PrAS, which includes an enormous amount of proteome data (over 500,000 protein sequences of 34 species in total) and is available via the web interface at http://alga-pras.riken.jp/. To obtain protein information from the Alga-PrAS database, four search functions—Property Search, Identifier (ID) Search, Keyword Search and Sequence Search—are provided in the Alga-PrAS database. These are detailed below.
Property Search
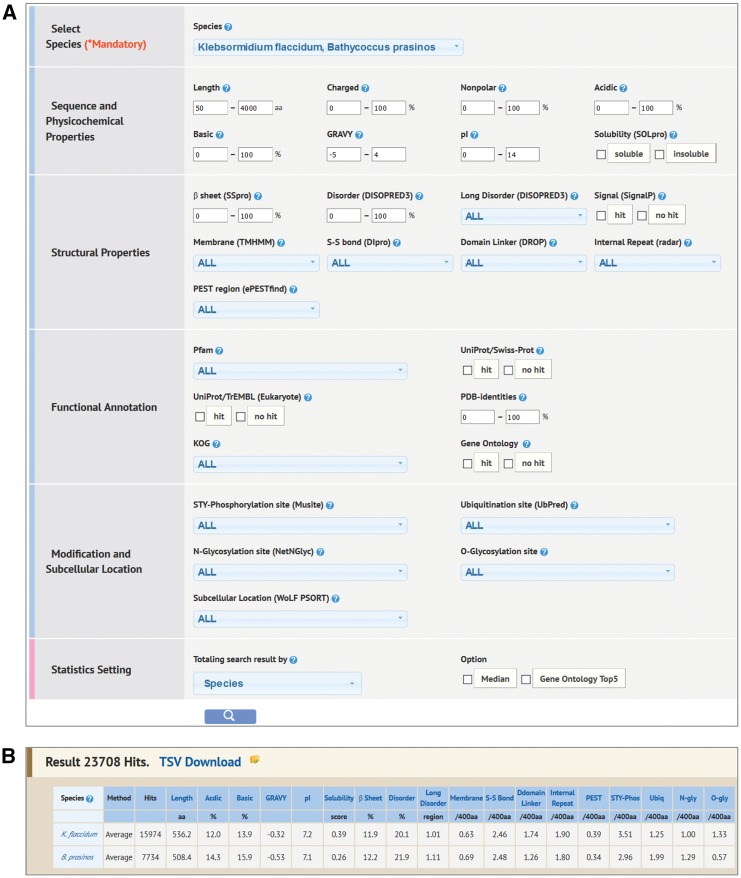
Property Search is the most comprehensive search function for accessing Alga-PrAS data. It provides a search function from 28 protein properties against 34 species proteomes (Fig. 1A). On the results page, a summary of the searched data containing average or median values for each property is shown in a summary statistics table (Fig. 1B). Subsequently, when users click on one of the hyperlinked items (e.g. species, taxonomic class) on the left side of the table, IDs belonging to the selected items are listed on the same page. The listed IDs are linked to the annotation detail page of each protein (Fig. 2). In this search there is also a convenient function for comparison analysis among the Alga-PrAS data. By setting the display option, the summary statistics table can be sorted by species, taxonomic classification, habitat, unicellularity or multicellularity, protein cluster and KOG, meaning that biological species can be selected by users based on common classification terms (land plants, green algae, red algae, Glaucophyceae, oomycetes, diatoms and other microalgae), habitat (freshwater, marine, terrestrial and ubiquitous), whether an organism is composed of one or multiple cells, species-specific or common protein clusters by orthologous clustering with the OrthoMCL tool (single-species cluster, all-species cluster and other) (Fischer et al. 2011), or 25 KOG function categories (Koonin et al. 2004) (Supplementary Tables S1 and S3). In addition, to visualize numeric data the user can click a property item in the summary statistics table and display a bar chart frame.
Fig. 1.
Property Search interface. (A) Users can search by multiple protein properties on the Property Search page. (B) Example of a summary table from the Property Search results.
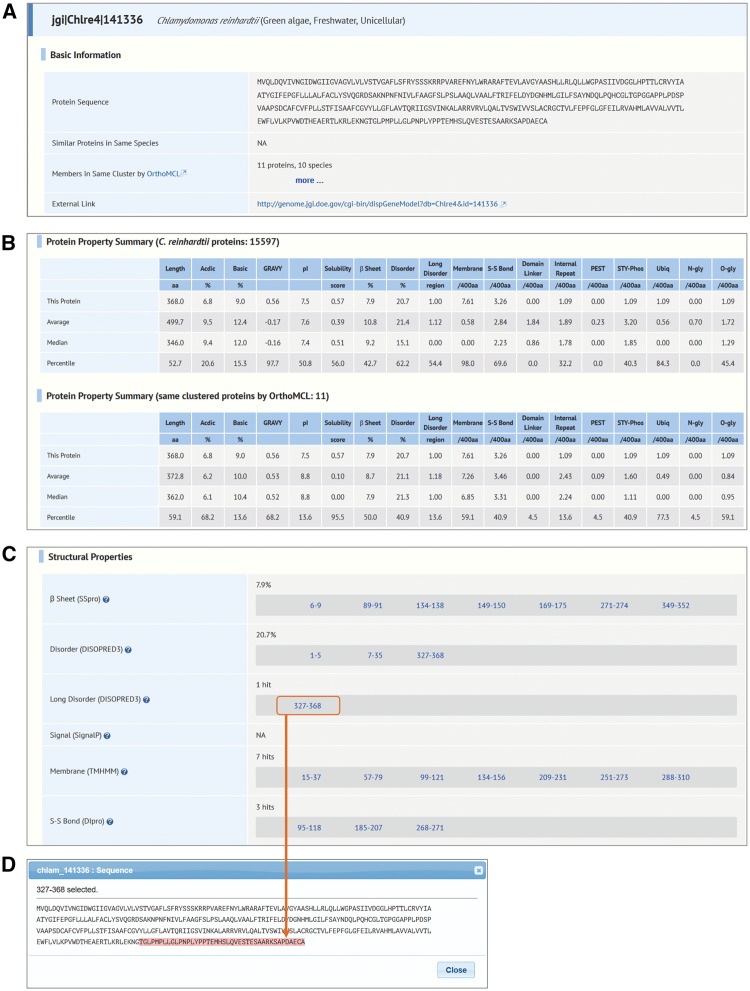
Fig. 2.
Typical examples of annotation detail page. (A) Basic information on a protein in Alga-PrAS. (B) Summary with average, median and percentile values in relation to proteins from identical species (upper portion) and identical clustered proteins by OrthoMCL (lower portion). (C) Structural properties. (D) Sequence window for highlighting position data for regions or sites.
ID Search and Keyword Search
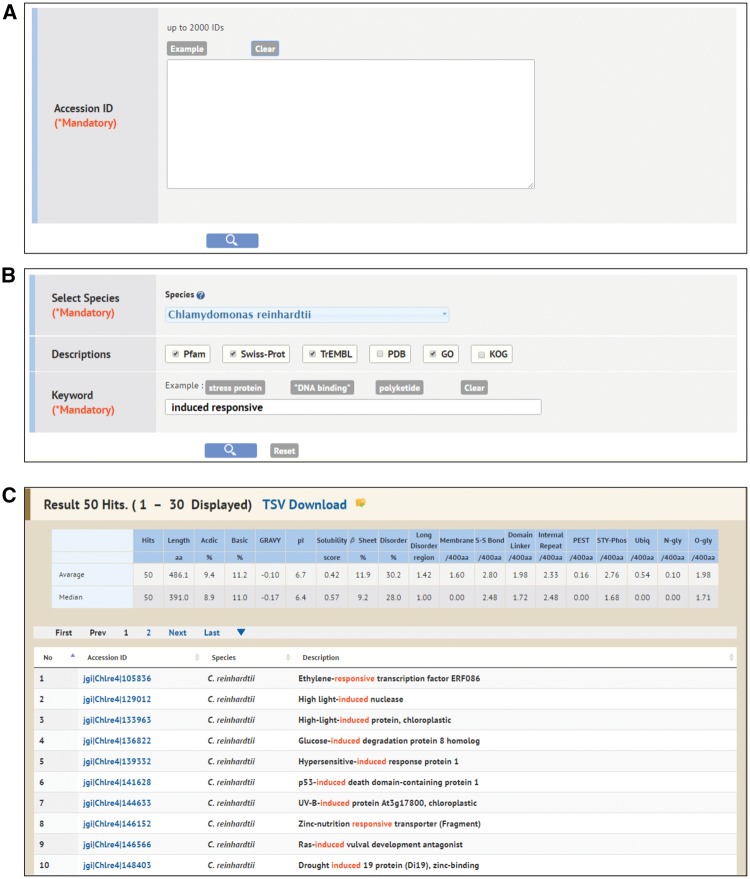
ID Search is a simpler search function for accessing the Alga-PrAS data if the user knows the accession IDs of proteins on public protein databases such as UniProtKB and Pfam. It provides a search function by inputting arbitrary IDs in the text box as a query (Fig. 3A). Keyword Search is an annotation search function against the assigned descriptions of the Pfam, UniProt/Swiss-Prot, UniProt/TrEMBL, PDB, GO and KOG databases housed in advance in Alga-PrAS (Fig. 3B). A multiple keyword search is performed when the introduced keywords are separated by spaces. In addition, an exact phrase search is performed by enclosing keywords within quotation marks, and, to exclude specific words, users can use a hyphen as a prefix for the keyword they wish to exclude. For example, using (Myb -like) as a search keyword excludes the word ‘like’ from the search results. The ID list from the ID or Keyword Search is shown on the results page (Fig. 3C). Listed IDs are linked to the annotation detail page of each protein in the same manner as that of Property Search (Fig. 2).
Fig. 3.
Interfaces of ID Search and Keyword Search. (A) ID Search. This provides a search function by inputting arbitrary IDs in the text box as a query. (B) Keyword Search. This is an annotation search function against the assigned descriptions of the public databases. (C) Example of the results of Keyword Search. The example is the search result for the species Chlamydomonas reinhardtii, the description Pfam, Swiss-Prot and TrEMBL, and the keywords induced responsive.
Sequence Search
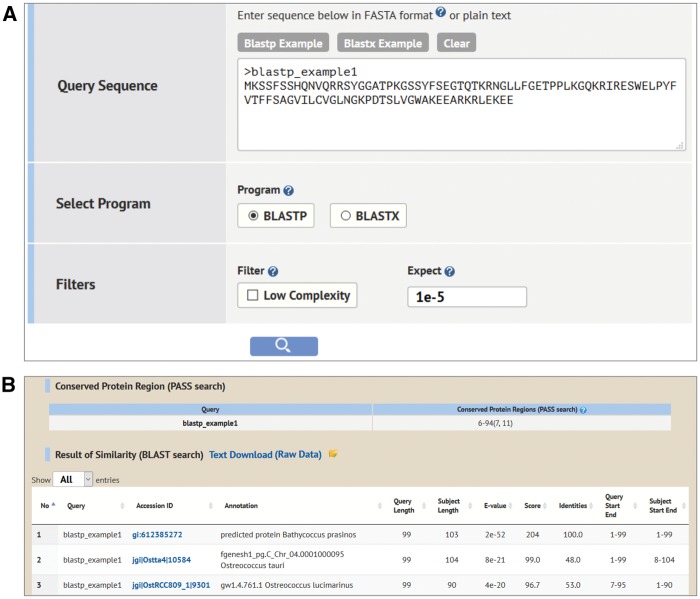
Sequence Search contains two search processes for algal data with users’ arbitrary sequences (Fig. 4A). One is a BLAST (Altschul et al. 1997, Altschul et al. 2005) search against the algal sequences in Alga-PrAS. The other is a conserved protein region search with the PASS tool (Kuroda et al. 2000), which determines the N-terminal site and the C-terminal site of conserved protein regions among diverse organisms using the BLAST result. Therefore, users can confirm the information on sequence similarity and conserved protein regions between their arbitrary sequences and the algal sequences housed in Alga-PrAS. This search allows protein or nucleic acid sequences to be submitted in the FASTA format as a query, with the option of a cutoff e-value. The result tables for BLAST and PASS searches are shown in the footer of the same page (Fig. 4B). The searched IDs are linked to the annotation detail page of each protein in the same manner as for the results page of the search functions mentioned above (Fig. 2).
Fig. 4.
Sequence Search interface. (A) Sequence Search allows protein or nucleic acid sequences to be submitted in the FASTA format as a query with the option of a cutoff e-value. (B) Example of Sequence Search results. The result tables for BLAST and PASS searches show that the conserved protein region is located from six to 94 amino acids of the query protein sequence.
Annotation detail page
The annotation detail page displays all the information available for an individual protein. The basic information, including amino acid sequence, IDs of similar proteins omitted in the clustering process by the CD-HIT tool in order to remove redundant sequences, and the IDs in the same cluster of all protein sequences in Alga-PrAS by the OrthoMCL tool, is displayed in the top part of page (Fig. 2A). Next, the summary tables of protein properties for proteomes of identical species and clustered proteins are displayed under the basic information section (Fig. 2B). Items in the summary tables consist of average and median values and percentile ranks for each protein property. Thus, the status of the query protein can be easily recognized among the Alga-PrAS data. Finally, all protein properties from sequence analyses are displayed under the summary tables (Fig. 2C). When users click the hyperlinked position data for regions or sites, these are highlighted on the protein sequence in an additional window (Fig. 2D). Additionally, external links to protein sequences and annotations (Pfam, UniProtKB, PDB, KOG and GO databases) are provided to enable verification with the original information on the resource websites.
Download
Users can download all information at the resources page in Alga-PrAS (http://alga-pras.riken.jp/menta.cgi/algapras/resources). In addition to the bulk download page, search results can also be downloaded as a tab-separated value (TSV) file each time a search is performed.
Examples of utilization of Alga-PrAS
Exploring candidate G protein-coupled receptors (GPCRs)
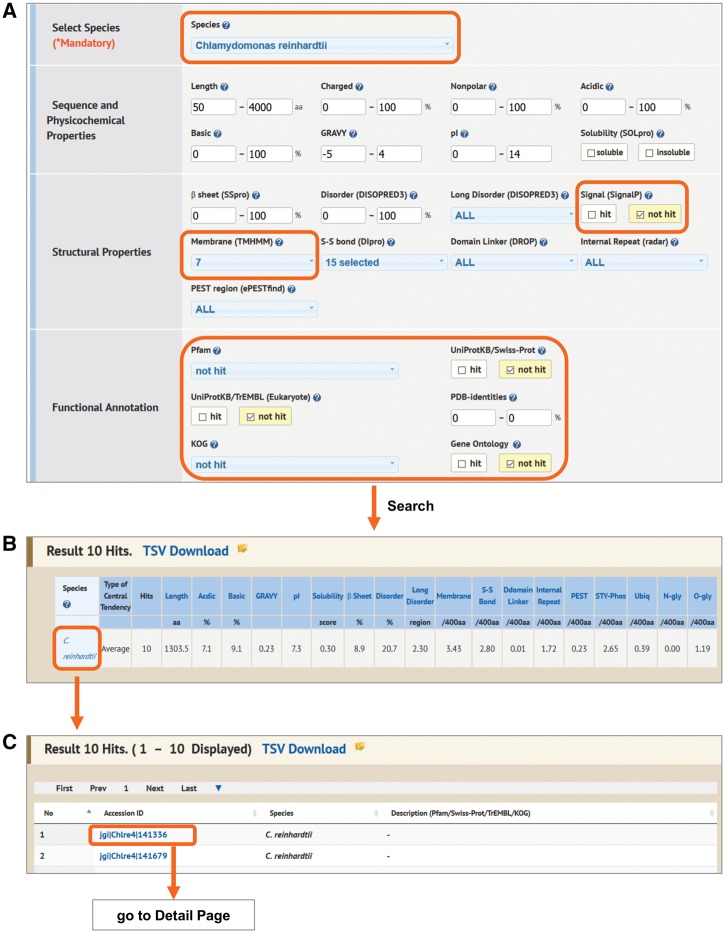
GPCRs constitute a large and diverse family of proteins that regulate various cellular functions involved in physiological responses (Guan et al. 1992, Pierce et al. 2002). We explored GPCR candidates in C. reinhardtii protein sequences known to contain seven membrane helix domain receptors and to lack a cleavable signal sequence (Singer 1990, Guan et al. 1992). First, we set ‘Chlamydomonas reinhardtii’ in the Species field (e.g. ‘7’ in Membrane, ‘not hit’ in Signal, Pfam, UniProtKB/Swiss-Prot, UniProtKB/TrEMBL, KOG and Gene Ontology, and ‘0%’ in PDB on the Property Search page; Fig. 5A). Negative settings in Pfam to PDB were intended to retrieve proteins that do not have functional annotations in these databases. This approach identified 10 protein sequences as candidate GPCRs (Fig. 5B). Next, we click on ‘C. reinhardtii’ in the Species column on the summary table; the accession IDs retrieved as a result of the above search process are displayed (Fig. 5C). When one of the protein IDs (e.g. jgi|Chlre4|141336) is clicked, the annotation detail page of the protein is displayed (Fig. 2). In this page the following information is shown: (i) other proteins belonging to the same cluster in the ‘Members in same cluster by OrthoMCL’ field in basic information (Fig. 2A), and (ii) the summary statistics of protein properties in the C. reinhardtii proteome and of the members of the same cluster (Fig. 2B) in the protein properties and the structural properties (Fig. 2C).
Fig. 5.
Search example of the exploration of candidates of G protein-coupled receptors (GPCRs). The settings for Property Search are as follows; ‘Chlamydomonas reinhardtii’ in the Species field (e.g. ‘7’ in Membrane), ‘not hit’ in Signal, Pfam, UniProtKB/Swiss-Prot, UniProtKB/TrEMBL, KOG and Gene Ontology, and ‘0%’ in PDB (A). The results identified 10 protein sequences as candidate GPCRs (B). Users click ‘C. reinhardtii’ on the Species column on the summary table, and the accession IDs which are searched by the above process are then displayed (C).
Number of PTMs in species-specific and common protein clusters in proteomes of land plants and algae
It is reported that the conservation of protein structure regions has been associated with higher amino acid substitution rates and faster evolution (Kim et al. 2008, Mosca et al. 2012, Brunquell et al. 2014). Thus, differences in the number of PTMs between species-specific protein clusters and common protein clusters of algae proteomes may be expected. To explore this in Alga-PrAS, protein clusters for all the proteins used in this study were created using the OrthoMCL tool (Fischer et al. 2011), housing in advance the results in the Alga-PrAS database as described previously. Then, protein clusters consisting of all 34 species used in this study were defined as common protein clusters. Protein clusters involving only one species were regarded as species-specific protein clusters. The content of all PTM parameters, including phosphorylation, glycosylation and ubiquitination in species-specific protein clusters or in common protein clusters of each taxonomic class, is shown in Table 3. In this analysis, we normalized the number of PTM sites to the same length (400 amino acids) based on the dataset’s average protein length. Information regarding PTM parameters can be obtained from the bulk download file. The contents of phosphorylation parameters were 1.1–5.4 times higher in species-specific protein clusters than in common protein clusters and the occurrence of phosphorylation in ratios of species-specific/common protein clusters in algal species was higher than in land plants (Table 3). This result may imply that algal species, which are simpler than land plants, utilize phosphorylation better than land plants. To date, many studies have been conducted on plant protein phosphorylation sites on photosynthetic membranes, and under a variety of conditions from biotic and abiotic stresses to changing nutrient environments. The principle of activation and inactivation of proteins by phosphorylation and the function of phosphorylated amino acid residues as docking sites have also been well characterized in the field of plant signal transduction (Turkina et al. 2006, Turkina and Vener 2007, Camoni et al. 2000, Nakagami et al. 2010).
Table 3.
Preference of protein disorder and PTMs in species-specific protein clusters and common protein clusters for each taxonomic class
| Taxonomic class | Disorder | S-pho/400aad | T-pho/400aad | Y-pho/400aad | O-gly/400aad | N-gly/400aad | Ubi/400aad | |
|---|---|---|---|---|---|---|---|---|
| Land plants | Specifica | 16% | 1.3 | 0.5 | 0.5 | 0.9 | 1.3 | 1.1 |
| Commonb | 13% | 0.7 | 0.3 | 0.4 | 0.6 | 1.2 | 0.7 | |
| S/C ratioc | 1.2 | 2.0 | 1.7 | 1.1 | 1.4 | 1.1 | 1.5 | |
| Green algae | Specific | 20% | 2.4 | 1.2 | 0.6 | 1.8 | 0.9 | 0.9 |
| Common | 12% | 0.6 | 0.3 | 0.5 | 0.8 | 0.9 | 0.6 | |
| S/C ratio | 1.7 | 4.0 | 3.6 | 1.4 | 2.4 | 0.9 | 1.6 | |
| Red algae | Specific | 12% | 1.7 | 0.9 | 0.6 | 1.4 | 1.0 | 0.9 |
| Common | 14% | 0.7 | 0.4 | 0.5 | 0.8 | 1.0 | 0.6 | |
| S/C ratio | 0.9 | 2.3 | 2.1 | 1.3 | 1.7 | 1.0 | 1.5 | |
| Glaucophyceae | Specific | 14% | 2.3 | 1.0 | 0.5 | 1.8 | 0.8 | 0.8 |
| Common | 10% | 0.5 | 0.3 | 0.3 | 0.9 | 1.0 | 0.6 | |
| S/C ratio | 1.4 | 4.9 | 3.6 | 1.5 | 2.0 | 0.8 | 1.4 | |
| Oomycetes | Specific | 14% | 1.3 | 0.7 | 0.6 | 0.9 | 1.3 | 0.8 |
| Common | 12% | 0.6 | 0.3 | 0.4 | 0.7 | 1.1 | 0.7 | |
| S/C ratio | 1.1 | 2.3 | 2.2 | 1.3 | 1.4 | 1.1 | 1.2 | |
| Diatoms | Specific | 20% | 1.8 | 0.8 | 0.6 | 1.0 | 2.1 | 1.8 |
| Common | 10% | 0.3 | 0.2 | 0.4 | 0.6 | 1.2 | 0.7 | |
| S/C ratio | 2.0 | 5.4 | 4.7 | 1.7 | 1.9 | 1.7 | 2.7 | |
| Other microalgae | Specific | 16% | 2.1 | 0.9 | 0.6 | 1.2 | 1.0 | 1.4 |
| Common | 11% | 0.7 | 0.4 | 0.4 | 0.8 | 0.9 | 0.7 | |
| S/C ratio | 1.4 | 3.1 | 2.7 | 1.4 | 1.6 | 1.1 | 2.1 |
a The Specific category (species-specific protein clusters) involves just one species in a cluster using the OrthoMCL tool.
b The Common category (common protein clusters) involves all 34 species used in this study.
c Ratio of specific to common values.
d Average of normalized value of predicted PTM sites. The number of predicted PTM sites was normalized per 400 amino acids (aa).
Conclusion
Alga-PrAS is the most comprehensive resource for integrating abundant algal proteome information, and has an effective interface to enable the interpretation of algal proteome features. Importantly, the system can be expected to enhance gene functional annotation and further developments in algal species.
Materials and Methods
Resources for protein sequences
In this study we used 31 algal proteome sequence sets involving 12 green algae, five red algae, one Glaucophyceae, four oomycetes, three diatoms and six other algal species (Table 4). Three land plant species, Arabidopsis thaliana (Swarbreck et al. 2008), Selaginella moellendorffii (Banks et al. 2011) and Physcomitrella patens (Rensing et al. 2008) were also used (Table 4). Non-redundant protein sequence sets were prepared. First, sequences of less than 50 and more than 4,000 amino acids were excluded to avoid calculation failure in the prediction processes performed with DIpro (Cheng et al. 2006), SSpro (Cheng et al. 2005) and DROP (Ebina et al. 2011). To prepare non-redundant proteome sequence sets of each species, individual protein clusters of each species were created with the CD-HIT program (Fu et al. 2012) with default runtime parameters, and a protein sequence set specific to each species was used as input data..2
Table 4.
List of protein sequence resources in this study
| Classification | Species | Proteome resources | References for genomic analysis |
|---|---|---|---|
| Green algae | Klebsormidium flaccidum | Klebsormidium flaccidum Genome Projectg | Hori et al. 2014 |
| Ostreococcus lucimarinus | JGI Genome Portalh | Palenik et al. 2007 | |
| Ostreococcus tauri | JGI Genome Portalh | Derelle, et al. 2006 | |
| Micromonas pusilla | JGI Genome Portalh | Worden et al. 2009 | |
| Micromonas sp. RCC299 | JGI Genome Portalh | Worden, et al. 2009 | |
| Bathycoccus prasinos | NCBIi | Moreau et al. 2012 | |
| Volvox carteri | JGI Genome Portalh | Prochnik et al. 2010 | |
| Chlamydomonas reinhardtii | JGI Genome Portalh | Merchant, et al. 2007 | |
| Monoraphidium neglectum | NCBIi | Bogen et al. 2013 | |
| Coccomyxa subellipsoidea | JGI Genome Portalh | Blanc et al. 2010 | |
| Chlorella variabilis | JGI Genome Portalh | Blanc, et al. 2010 | |
| Auxenochlorella protothecoides | NCBIi | Gao et al. 2014 | |
| Red algae | Cyanidioschyzon merolae | Cyanidioschyzon merolae Genome Projectj | Matsuzaki, et al. 2004, Nozaki et al. 2007 |
| Galdieria sulphuraria | NCBIi | Schonknecht, et al. 2013 | |
| Pyropia yezoensis | NRIFSk | Nakamura et al. 2013 | |
| Chondrus crispus | NCBIi | Collen et al. 2013 | |
| Porphyridium purpureum | Porphyridium purpureum Genome Projectl | Bhattacharya et al. 2013 | |
| Glaucophyceae | Cyanophora paradoxa | Cyanophora Genome Projectm | Price et al. 2012 |
| Oomycetes | Phytophthora ramorum | JGI Genome Portalh | Tyler et al. 2006 |
| Phytophthora sojae | JGI Genome Portalh | Tyler et al. 2006 | |
| Phytophthora infestans | Superfamily databasen | Haas et al. 2009 | |
| Phytophthora capsici | JGI Genome Portalh | Lamour et al. 2012 | |
| Diatoms | Phaeodactylum tricornutum | JGI Genome Portalh | Bowler et al. 2008 |
| Fragilariopsis cylindrus sp. CCMP1102 | JGI Genome Portalh | http://genome.jgi.doe.gov/Fracy1/Fracy1.info.html | |
| Thalassiosira pseudonana | JGI Genome Portalh | Armbrust et al. 2004 | |
| Other algal species | Aureococcus anophagefferensa | JGI Genome Portalh | Gobler et al. 2011 |
| Ectocarpus siliculosusb | JGI Genome Portalh | Cock et al. 2010 | |
| Symbiodinium minutumc | OISTo | Shoguchi et al. 2013 | |
| Emiliania huxleyid | NCBIi | Read et al. 2013 | |
| Guillardia thetae | NCBIi | Curtis et al. 2012 | |
| Bigelowiella natansf | JGI Genome Portalh | Curtis et al. 2012 | |
| Land plants | Arabidopsis thaliana | TAIRp | Swarbreck et al. 2008 |
| Selaginella moellendorffii | JGI Genome Portalh | Banks et al. 2011 | |
| Physcomitrella patens | JGI Genome Portalh | Rensing et al. 2008 |
a–f Other algal species (Aureococcus anophagefferens, Ectocarpus siliculosus, Symbiodinium minutum, Emiliania huxleyi, Guillardia theta and Bigelowiella natans) belong to Pelagophyceae, Phaeophyceae, Dinophyceae, Haptophyceae, Cryptophyceae and Chlorarachniophyceae, respectively.
Calculation of protein properties
Physicochemical properties
The percentages of acidic, basic, charged and non-polar amino acids, as well as protein length and isoelectric point (pI), were calculated using the ProteoMix tool (Chikayama et al. 2004). The GRAVY index was calculated with the GRAVY algorithm (Kyte and Doolittle 1982). Protein solubility was determined using the SOLpro tool (Magnan et al. 2009).
Secondary structural properties
To detect protein properties related to secondary structure, we used the following tools: SignalP4.0 (Petersen et al. 2011), TMHMM2.0 (Krogh et al. 2001), DROP (Ebina et al. 2011), DIpro2.0 (Cheng et al. 2006), SSpro4 (Cheng et al. 2005), RADAR (Heger and Holm 2000), DISOPRED3 (Jones and Cozzetto 2015) and ePESTfind of EMBOSS (Rogers et al. 1986, Rice et al. 2000) to determine the presence of signal peptides, transmembrane helix domains, interdomain linkers, S–S bonds, secondary structures, internal repeats, intrinsically disordered regions and PEST regions, respectively.
Functional and structural annotations
We assigned protein annotations of KOG (Tatusov et al. 2000), UniProt/Swiss-Prot (Boutet et al. 2016), UniProtKB/TrEMBL (eukaryote) (Bateman et al. 2015) and PDB (Westbrook et al. 2003, Berman et al. 2014) using the BLASTP program with an e-value lower than 1e−10. The Pfam (Finn et al. 2016) and GO terms (Blake et al. 2015) were detected using InterProScan5 software (Hunter et al. 2012).
Modification and subcellular localization
To infer PTM and subcellular localization, we used the following tools and algorithms. Serine (Ser; S), threonine (Thr; T) or tyrosine (Tyr; Y) phosphorylation sites were detected with Musite1.0.1 (Gao et al. 2010) with the database option of Eukaryote-General-Ser-Thr;Eukaryote-General-Tyr. O-glycosylation sites were detected based on Gomord’s algorithm (Gomord et al. 2010). N-glycosylation sites were detected by combining the results of the NetNglyc1.0 tool (http://www.cbs.dtu.dk/services/NetNGlyc) with the signal peptide (SignalP) option and the TMHMM2.0 tool. Thus, we detected extracellular N-glycosylation sites with TMHMM2.0, and the number of signal peptides in the sequence was calculated with SignalP from NetNglyc1.0 to remove false-positive data with NetNglyc1.0. Ubiquitination sites were detected with the UbPred tool (Radivojac et al. 2010) with a medium confidence option. Transmembrane helix regions were detected with the TMHMM2.0 tool. Subcellular localizations were detected with the WoLF PSORT tool (Horton et al. 2007). Additionally, for the protein sequences of the diatoms Fragilariopsis cylindrus (CCMP 1102), Phaeodactylum tricornutum and Thalassiosira pseudonana, the cryptophyte Guillardia theta and the dinoflagellate Symbiodinium minutum, we used the HECTAR tool (Gschloessl et al. 2008) because the chloroplasts of these five algal species evolved from secondary endosymbiosis (Gruber et al. 2015).
Classification of species-specific and common protein clusters
To determine the number of PTMs in species-specific and common protein clusters in proteomes of land plants and algae, we created protein clusters among all the protein sequences in this study. First, we calculated pairwise sequence similarities between all the protein sequences by using the BLASTP program with an e-value lower than 1e−5. Subsequently, protein clusters were estimated by the Markov Clustering (MCL) algorithm employed in OrthoMCL1.4 (Fischer et al. 2011) with the BLASTP results and the default runtime parameters. Finally, a singlet and a cluster consisting of only one species were classified as a species-specific protein, and a cluster consisting of all 34 species was classified as a common protein cluster.
System availability and implementation
Alga-PrAS was implemented in the Linux operating system (CentOS 6.8, 64 bit) with a MENTA web application framework based on Perl 5.1.0 and MySQL 5.7.13 as a database engine, and tested on the following web browsers: Microsoft Edge 25, Internet Explorer 10+, Google Chrome 51+ and Firefox 41+.
Supplementary Data
Supplementary data are available at PCP online.
Supplementary Material
Acknowledgments
We thank Takuhiro Yoshida, Hiroaki Tanabe (RIKEN) and Alexander A. Tokmakov (Kyoto Sangyo University) for their helpful comments on analyses and computational support.
Glossary
Abbreviations
- Alga-PrAS
Algal Protein Annotation Suite
- BLAST
Basic Local Alignment Search Tool
- GRAVY
grand average value of hydropathicity index
- IDR
intrinsically disordered region
- NCBI
National Center for Biotechnology Information
- PTM
post-translational modification
Funding
This work was partly supported by a Grant-in-Aid for Young Scientists (B) (18700106 to T.S.) from the Japan Society for the Promotion of Science.
References
- Akiyama K., Kurotani A., Iida K., Kuromori T., Shinozaki K., Sakurai T. (2014) RARGE II: an integrated phenotype database of Arabidopsis mutant traits using a controlled vocabulary. Plant Cell Physiol. 55: E4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S.F., Madden T.L., Schaffer A.A., Zhang J.H., Zhang Z., Miller W., et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25: 3389–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S.F., Wootton J.C., Gertz E.M., Agarwala R., Morgulis A., Schaffer A.A., et al. (2005) Protein database searches using compositionally adjusted substitution matrices. FEBS Journal 272: 5101–5109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aoki Y., Okamura Y., Ohta H., Kinoshita K., Obayashi T. (2016a) ALCOdb: Gene Coexpression Database for Microalgae. Plant Cell Physiol. 57: E3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aoki Y., Okamura Y., Tadaka S., Kinoshita K., Obayashi T. (2016b) ATTED-II in 2016: a plant coexpression database towards lineage-specific coexpression. Plant Cell Physiol. 57: E5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armbrust E.V., Berges J.A., Bowler C., Green B.R., Martinez D., Putnam N.H., et al. (2004) The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306: 79–86. [DOI] [PubMed] [Google Scholar]
- Banks J.A., Nishiyama T., Hasebe M., Bowman J.L., Gribskov M., dePamphilis C., et al. (2011) The Selaginella genome identifies genetic changes associated with the evolution of vascular plants. Science 332: 960–963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bateman A., Martin M.J., O'Donovan C., Magrane M., Apweiler R., Alpi E., et al. (2015) UniProt: a hub for protein information. Nucleic Acids Res. 43: D204–D212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beheshtipour H., Mortazavian A.M., Mohammadi R., Sohrabvandi S., Khosravi-Darani K. (2013) Supplementation of Spirulina platensis and Chlorella vulgaris algae into probiotic fermented milks. Compr. Rev. Food Sci. Food Saf. 12: 144–154. [Google Scholar]
- Berardini T.Z., Reiser L., Li D.H., Mezheritsky Y., Muller R., Strait E., et al. (2015) The arabidopsis information resource: making and mining the "gold standard" annotated reference plant genome. Genesis 53: 474–485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berman H.M., Kleywegt G.J., Nakamura H., Markley J.L. (2014) The Protein Data Bank archive as an open data resource. J. Comput. Aided Mol. Des. 28: 1009–1014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharya D., Price D.C., Chan C.X., Qiu H., Rose N., Ball S., et al. (2013) Genome of the red alga Porphyridium purpureum. Nat. Commun. 4: 1941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharya D., Qiu H., Price D.C. (2015) Why we need more algal genomes. J. Phycol. 51: 1–5. [DOI] [PubMed] [Google Scholar]
- Blaby I.K., Blaby-Haas C.E., Tourasse N., Hom E.F.Y., Lopez D., Aksoy M., et al. (2014) The Chlamydomonas genome project: a decade on. Trends Plant Sci. 19: 672–680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blake J.A., Christie K.R., Dolan M.E., Drabkin H.J., Hill D.P., Ni L., et al. (2015) Gene Ontology Consortium: going forward. Nucleic Acids Res. 43: D1049–D1056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanc G., Duncan G., Agarkova I., Borodovsky M., Gurnon J., Kuo A., et al. (2010) The Chlorella variabilis NC64A genome reveals adaptation to photosymbiosis, coevolution with viruses, and cryptic sex. Plant Cell 22: 2943–2955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogen C., Al-Dilaimi A., Albersmeier A., Wichmann J., Grundmann M., Rupp O., et al. (2013) Reconstruction of the lipid metabolism for the microalga Monoraphidium neglectum from its genome sequence reveals characteristics suitable for biofuel production. BMC Genomics 14: 926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borowitzka M.A. (2013) High-value products from microalgae—their development and commercialisation. J. Appl. Phycol. 25: 743–756. [Google Scholar]
- Boutet E., Lieberherr D., Tognolli M., Schneider M., Bansal P., Bridge A.J., et al. (2016) UniProtKB/Swiss-Prot, the manually annotated section of the UniProt KnowledgeBase: how to use the entry view. Methods Mol.Biol. 1374: 23–54. [DOI] [PubMed] [Google Scholar]
- Bowler C., Allen A.E., Badger J.H., Grimwood J., Jabbari K., Kuo A., et al. (2008) The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456: 239–244. [DOI] [PubMed] [Google Scholar]
- Brunquell J., Yuan J., Erwin A., Westerheide S.D., Xue B. (2014) DBC1/CCAR2 and CCAR1 are largely disordered proteins that have evolved from one common ancestor. Biomed Res. Int. 2014: 418458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camoni L., Iori V., Marra M., Aducci P. (2000) Phosphorylation-dependent interaction between plant plasma membrane H+-ATPase and 14‐3‐3 proteins. J. Biol. Chem. 275: 9919–9923. [DOI] [PubMed] [Google Scholar]
- Carvalho A.P., Meireles L.A., Malcata F.X. (2006) Microalgal reactors: a review of enclosed system designs and performances. Biotechnol. Prog. 22: 1490–1506. [DOI] [PubMed] [Google Scholar]
- Cheng J., Randall A.Z., Sweredoski M.J., Baldi P. (2005) SCRATCH: a protein structure and structural feature prediction server. Nucleic Acids Res. 33: W72–W76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng J.L., Saigo H., Baldi P. (2006) Large-scale prediction of disulphide bridges using kernel methods, two-dimensional recursive neural networks, and weighted graph matching. Proteins 62: 617–629. [DOI] [PubMed] [Google Scholar]
- Chikayama E., Kurotani A., Kuroda Y., Yokoyama S. (2004) ProteoMix: an integrated and flexible system for interactively analyzing large numbers of protein sequences. Bioinformatics 20: 2836–2838. [DOI] [PubMed] [Google Scholar]
- Chisti Y. (2007) Biodiesel from microalgae. Biotechnol. Adv. 25: 294–306. [DOI] [PubMed] [Google Scholar]
- Clemente H.S., Jamet E. (2015) WallProtDB, a database resource for plant cell wall proteomics. Plant Methods 11(1): 2 doi: 10.1186/s13007‐015‐0045-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cock J.M., Sterck L., Rouze P., Scornet D., Allen A.E., Amoutzias G., et al. (2010) The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature 465: 617–621. [DOI] [PubMed] [Google Scholar]
- Collen J., Porcel B., Carre W., Ball S.G., Chaparro C., Tonon T., et al. (2013) Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proc. Natl Acad. Sci. USA 110: 5247–5252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Curtis B.A., Tanifuji G., Burki F., Gruber A., Irimia M., Maruyama S., et al. (2012) Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature 492: 59–65. [DOI] [PubMed] [Google Scholar]
- Derelle E., Ferraz C., Rombauts S., Rouze P., Worden A.Z., Robbens S., et al. (2006) Genome analysis of the smallest free-living eukaryote Ostreococcus tauri unveils many unique features. Proc. Natl Acad. Sci. USA 103: 11647–11652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong Q.F., Schlueter S.D., Brendel V. (2004) PlantGDB, plant genome database and analysis tools. Nucleic Acids Res. 32: D354–D359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Draaisma R.B., Wijffels R.H., Slegers P.M., Brentner L.B., Roy A., Barbosa M.J. (2013) Food commodities from microalgae. Curr. Opin. Biotechnol. 24: 169–177. [DOI] [PubMed] [Google Scholar]
- Ebina T., Toh H., Kuroda Y. (2011) DROP: an SVM domain linker predictor trained with optimal features selected by random forest. Bioinformatics 27: 487–494. [DOI] [PubMed] [Google Scholar]
- Finn R.D., Coggill P., Eberhardt R.Y., Eddy S.R., Mistry J., Mitchell A.L., et al. (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44: D279–D285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fischer S., Brunk B.P., Chen F., Gao X., Harb O.S., Iodice J.B., et al. (2011) Using OrthoMCL to assign proteins to OrthoMCL-DB groups or to cluster proteomes into new ortholog groups. Curr. Protoc. Bioinform. 35: 6.12.1–6.12.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu L., Niu B., Zhu Z., Wu S., Li W. (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28: 3150–3152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao C.F., Wang Y., Shen Y., Yan D., He X., Dai J.B., et al. (2014) Oil accumulation mechanisms of the oleaginous microalga Chlorella protothecoides revealed through its genome, transcriptomes, and proteomes. BMC Genomics 15: 582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao J.J., Thelen J.J., Dunker A.K., Xu D. (2010) Musite, a tool for global prediction of general and kinase-specific phosphorylation sites. Mol. Cell. Proteomics 9: 2586–2600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gobler C.J., Berry D.L., Dyhrman S.T., Wilhelm S.W., Salamov A., Lobanov A.V., et al. (2011) Niche of harmful alga Aureococcus anophagefferens revealed through ecogenomics. Proc. Natl Acad. Sci. USA 108: 4352–4357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomord V., Fitchette A.C., Menu-Bouaouiche L., Saint-Jore-Dupas C., Plasson C., Michaud D., et al. (2010) Plant-specific glycosylation patterns in the context of therapeutic protein production. Plant Biotechnol. J. 8: 564–587. [DOI] [PubMed] [Google Scholar]
- Goodstein D.M., Shu S.Q., Howson R., Neupane R., Hayes R.D., Fazo J., et al. (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40: D1178–D1186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruber A., Rocap G., Kroth P.G., Armbrust E.V., Mock T. (2015) Plastid proteome prediction for diatoms and other algae with secondary plastids of the red lineage. Plant J. 81: 519–528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gschloessl B., Guermeur Y., Cock J.M. (2008) HECTAR: a method to predict subcellular targeting in heterokonts. BMC Bioinform. 9: 393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan X.M., Kobilka T.S., Kobilka B.K. (1992) Enhancement of membrane insertion and function in a type iiib membrane-protein following introduction of a cleavable signal peptide. J. Biol. Chem. 267: 21995–21998. [PubMed] [Google Scholar]
- Guiry M.D., Guiry G.M., Morrison L., Rindi F., Valenzuela Miranda S., Mathieson A.C., et al. (2014) AlgaeBase: an on-line resource for algae. Cryptogam. Algol. 35: 105–115. [Google Scholar]
- Guo A.Y., Chen X., Gao G., Zhang H., Zhu Q.H., Liu X.C., et al. (2008) PlantTFDB: a comprehensive plant transcription factor database. Nucleic Acids Res. 36: D966–D969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haas B.J., Kamoun S., Zody M.C., Jiang R.H., Handsaker R.E., Cano L.M., et al. (2009) Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461: 393–398. [DOI] [PubMed] [Google Scholar]
- Heger A., Holm L. (2000) Rapid automatic detection and alignment of repeats in protein sequences. Proteins 41: 224–237. [DOI] [PubMed] [Google Scholar]
- Hori K., Maruyama F., Fujisawa T., Togashi T., Yamamoto N., Seo M., et al. (2014) Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation. Nat. Commun. 5: 3978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horton P., Park K.J., Obayashi T., Fujita N., Harada H., Adams-Collier C.J., et al. (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res. 35: W585–W587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter S., Jones P., Mitchell A., Apweiler R., Attwood T.K., Bateman A., et al. (2012) InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res. 40: D306–D312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imanishi T., Itoh T., Suzuki Y., O'Donovan C., Fukuchi S. (2004) Integrative annotation of 21,037 human genes validated by full-length cDNA. PLoS Biol 2(6): http://dx.doi.org/10.1371/journal.pbio.0020162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones D.T., Cozzetto D. (2015) DISOPRED3: precise disordered region predictions with annotated protein-binding activity. Bioinformatics 31: 857–863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keeling P.J., Burki F., Wilcox H.M., Allam B., Allen E.E., Amaral-Zettler L.A., et al. (2014) The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol 12(6): e1001889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim K.M., Park J.H., Bhattacharya D., Yoon H.S. (2014) Applications of next-generation sequencing to unravelling the evolutionary history of algae. Int. J. Syst. Evol. Microbiol. 64: 333–345. [DOI] [PubMed] [Google Scholar]
- Kim P.M., Sboner A., Xia Y., Gerstein M. (2008) The role of disorder in interaction networks: a structural analysis. Mol. Syst. Biol. 4: 179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koonin E.V., Fedorova N.D., Jackson J.D., Jacobs A.R., Krylov D.M., Makarova K.S., et al. (2004) A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5: R7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kourmpetis Y.A.I., van Dijk A.D.J., van Ham R.C.H.J., ter Braak C.J.F. (2011) Genome-wide computational function prediction of Arabidopsis proteins by integration of multiple data sources. Plant Physiol. 155: 271–281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogh A., Larsson B., von Heijne G., Sonnhammer E.L. (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305: 567–580. [DOI] [PubMed] [Google Scholar]
- Kudo T., Akiyama K., Kojima M., Makita N., Sakurai T., Sakakibara H. (2013) UniVIO: a multiple omics database with hormonome and transcriptome data from rice. Plant Cell Physiol. 54: E9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroda Y., Tani K., Matsuo Y., Yokoyama S. (2000) Automated search of natively folded protein fragments for high-throughput structure determination in structural genomics. Protein Sci. 9: 2313–2321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurotani A., Yamada Y., Shinozaki K., Kuroda Y., Sakurai T. (2015) Plant-PrAS: a database of physicochemical and structural properties and novel functional regions in plant proteomes. Plant Cell Physiol. 56: E11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R.F. (1982) A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157: 105–132. [DOI] [PubMed] [Google Scholar]
- Lamour K.H., Mudge J., Gobena D., Hurtado-Gonzales O.P., Schmutz J., Kuo A., et al. (2012) Genome sequencing and mapping reveal loss of heterozygosity as a mechanism for rapid adaptation in the vegetable pathogen Phytophthora capsici. Mol. Plant Microbe Interact. 25: 1350–1360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Magnan C.N., Randall A., Baldi P. (2009) SOLpro: accurate sequence-based prediction of protein solubility. Bioinformatics 25: 2200–2207. [DOI] [PubMed] [Google Scholar]
- Matsuzaki M., Misumi O., Shin I.T., Maruyama S., Takahara M., Miyagishima S.Y., et al. (2004) Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 428: 653–657. [DOI] [PubMed] [Google Scholar]
- May P., Christian J.O., Kempa S., Walther D. (2009) ChlamyCyc: an integrative systems biology database and web-portal for Chlamydomonas reinhardtii. BMC Genomics 10: 209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGarvey K.M., Goldfarb T., Cox E., Farrell C.M., Gupta T., Joardar V.S., et al. (2015) Mouse genome annotation by the RefSeq project. Mamm. Genome 26: 379–390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medipally S.R., Yusoff F.M., Banerjee S., Shariff M. (2015) Microalgae as sustainable renewable energy feedstock for biofuel production. Biomed Res. Int . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merchant S.S., Prochnik S.E., Vallon O., Harris E.H., Karpowicz S.J., Witman G.B., et al. (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245–250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreau H., Verhelst B., Couloux A., Derelle E., Rombauts S., Grimsley N., et al. (2012) Gene functionalities and genome structure in Bathycoccus prasinos reflect cellular specializations at the base of the green lineage. Genome Biol. 13: R74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mosca R., Pache R.A., Aloy P. (2012) The role of structural disorder in the rewiring of protein interactions through evolution. Mol. Cell. Proteomics 11: 014969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakagami H., Sugiyama N., Mochida K., Daudi A., Yoshida Y., Toyoda T., et al. (2010) Large-scale comparative phosphoproteomics identifies conserved phosphorylation sites in plants. Plant Physiol. 153: 1161–1174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura Y., Sasaki N., Kobayashi M., Ojima N., Yasuike M., Shigenobu Y., et al. (2013) The first symbiont-free genome sequence of marine red alga, Susabi-nori (Pyropia yezoensis). PLoS One 8: e57122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nordberg H., Cantor M., Dusheyko S., Hua S., Poliakov A., Shabalov I., et al. (2014) The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 42: D26–D31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nozaki H., Takano H., Misumi O., Terasawa K., Matsuzaki M., Maruyama S., et al. (2007) A 100%-complete sequence reveals unusually simple genomic features in the hot-spring red alga Cyanidioschyzon merolae. BMC Biol. 5: 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oates M.E., Stahlhacke J., Vavoulis D.V., Smithers B., Rackham O.J.L., Sardar A.J., et al. (2015) The SUPERFAMILY 1.75 database in 2014: a doubling of data. Nucleic Acids Res. 43: D227–D233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orlowski J., Bujnicki J.M. (2008) Structural and evolutionary classification of Type II restriction enzymes based on theoretical and experimental analyses. Nucleic Acids Res. 36: 3552–3569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ouyang S., Zhu W., Hamilton J., Lin H., Campbell M., Childs K., et al. (2007) The TIGR Rice Genome Annotation Resource: improvements and new features. Nucleic Acids Res 35: D883–D887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palenik B., Grimwood J., Aerts A., Rouze P., Salamov A., Putnam N., et al. (2007) The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proc. Natl Acad. Sci. USA 104: 7705–7710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Passell H., Dhaliwal H., Reno M., Wu B., Ben Amotz A., Ivry E., et al. (2013) Algae biodiesel life cycle assessment using current commercial data. J. Environ. Manag. 129: 103–111. [DOI] [PubMed] [Google Scholar]
- Petersen T.N., Brunak S., von Heijne G., Nielsen H. (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8: 785–786. [DOI] [PubMed] [Google Scholar]
- Pierce K.L., Premont R.T., Lefkowitz R.J. (2002) Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3: 639–650. [DOI] [PubMed] [Google Scholar]
- Price D.C., Chan C.X., Yoon H.S., Yang E.C., Qiu H., Weber A.P., et al. (2012) Cyanophora paradoxa genome elucidates origin of photosynthesis in algae and plants. Science 335: 843–847. [DOI] [PubMed] [Google Scholar]
- Prochnik S.E., Umen J., Nedelcu A.M., Hallmann A., Miller S.M., Nishii I., et al. (2010) Genomic analysis of organismal complexity in the multicellular green alga Volvox carteri. Science 329: 223–226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pruitt K.D., Tatusova T., Brown G.R., Maglott D.R. (2012) NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy. Nucleic Acids Res. 40: D130–D135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pruitt K.D., Tatusova T., Maglott D.R. (2007) NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 35: D61–D65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radivojac P., Vacic V., Haynes C., Cocklin R.R., Mohan A., Heyen J.W., et al. (2010) Identification, analysis, and prediction of protein ubiquitination sites. Proteins 78: 365–380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Read B.A., Kegel J., Klute M.J., Kuo A., Lefebvre S.C., Maumus F., et al. (2013) Pan genome of the phytoplankton Emiliania underpins its global distribution. Nature 499: 209–213. [DOI] [PubMed] [Google Scholar]
- Reijnders M.J.M.F., van Heck R.G.A., Lam C.M.C., Scaife M.A., dos Santos V.A.P.M., Smith A.G., et al. (2014) Green genes: bioinformatics and systems-biology innovations drive algal biotechnology. Trends Biotechnol. 32: 617–626. [DOI] [PubMed] [Google Scholar]
- Rensing S.A., Lang D., Zimmer A.D., Terry A., Salamov A., Shapiro H., et al. (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319: 64–69. [DOI] [PubMed] [Google Scholar]
- Rice P., Longden I., Bleasby A. (2000) EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 16: 276–277. [DOI] [PubMed] [Google Scholar]
- Rogers S., Wells R., Rechsteiner M. (1986) Amino acid sequences common to rapidly degraded proteins: the PEST hypothesis. Science 234: 364–368. [DOI] [PubMed] [Google Scholar]
- Saghatelianr A., Couso J.P. (2015) Discovery and characterization of smORF-encoded bioactive polypeptides. Nat. Chem. Biol 11: 909–916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakai H., Lee S.S., Tanaka T., Numa H., Kim J., Kawahara Y., et al. (2013) Rice Annotation Project Database (RAP-DB): an integrative and interactive database for rice genomics. Plant Cell Physiol. 54: E6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakurai T., Yamada Y., Sawada Y., Matsuda F., Akiyama K., Shinozaki K., et al. (2013) PRIMe Update: innovative content for plant metabolomics and integration of gene expression and metabolite accumulation. Plant Cell Physiol. 54: E5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schonknecht G., Chen W.H., Ternes C.M., Barbier G.G., Shrestha R.P., Stanke M., et al. (2013) Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote. Science 339: 1207–1210. [DOI] [PubMed] [Google Scholar]
- Shoguchi E., Shinzato C., Kawashima T., Gyoja F., Mungpakdee S., Koyanagi R., et al. (2013) Draft assembly of the Symbiodinium minutum nuclear genome reveals dinoflagellate gene structure. Curr. Biol. 23: 1399–1408. [DOI] [PubMed] [Google Scholar]
- Singer S.J. (1990) The structure and insertion of integral proteins in membranes. Annu. Rev. Cell Biol. 6: 247–296. [DOI] [PubMed] [Google Scholar]
- Sumiya N., Kawase Y., Hayakawa J., Matsuda M., Nakamura M., Era A., et al. (2015) Expression of cyanobacterial acyl-ACP reductase elevates the triacylglycerol level in the red alga Cyanidioschyzon merolae. Plant Cell Physiol. 56: 1962–1980. [DOI] [PubMed] [Google Scholar]
- Swarbreck D., Wilks C., Lamesch P., Berardini T.Z., Garcia-Hernandez M., Foerster H., et al. (2008) The Arabidopsis Information Resource (TAIR): gene structure and function annotation. Nucleic Acids Res 36: D1009–D1014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatusov R.L., Galperin M.Y., Natale D.A., Koonin E.V. (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28: 33–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torzillo G., Scoma A., Faraloni C., Giannelli L. (2015) Advances in the biotechnology of hydrogen production with the microalga Chlamydomonas reinhardtii. Crit. Rev. Biotechnol. 35: 485–496. [DOI] [PubMed] [Google Scholar]
- Turkina M.V., Kargul J., Blanco-Rivero A., Villarejo A., Barber J., Vener A.V. (2006) Environmentally modulated phosphoproteome of photosynthetic membranes in the green alga Chlamydomonas reinhardtii. Mol. Cell. Proteomics 5: 1412–1425. [DOI] [PubMed] [Google Scholar]
- Turkina M.V., Vener A.V. (2007) Identification of phosphorylated proteins. Methods Mol. Biol. 355: 305–316. [DOI] [PubMed] [Google Scholar]
- Tyler B.M., Tripathy S., Zhang X., Dehal P., Jiang R.H., Aerts A., et al. (2006) Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313: 1261–1266. [DOI] [PubMed] [Google Scholar]
- Vandepoele K., Van Bel M., Richard G., Van Landeghem S., Verhelst B., Moreau H., et al. (2013) pico-PLAZA, a genome database of microbial photosynthetic eukaryotes. Environ. Microbiol. 15: 2147–2153. [DOI] [PubMed] [Google Scholar]
- Westbrook J., Feng Z., Chen L., Yang H., Berman H.M. (2003) The Protein Data Bank and structural genomics. Nucleic Acids Res. 31: 489–491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wijffels R.H., Barbosa M.J. (2010) An outlook on microalgal biofuels. Science 329: 796–799. [DOI] [PubMed] [Google Scholar]
- Worden A.Z., Lee J.H., Mock T., Rouze P., Simmons M.P., Aerts A.L., et al. (2009) Green evolution and dynamic adaptations revealed by genomes of the marine picoeukaryotes Micromonas. Science 324: 268–272. [DOI] [PubMed] [Google Scholar]
- Yamamoto N., Kudo T., Fujiwara S., Takatsuka Y., Hirokawa Y., Tsuzuki M., et al. (2016) Pleurochrysome: a web database of Pleurochrysis transcripts and orthologs among heterogeneous algae. Plant Cell Physiol. 57: E6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youens-Clark K., Buckler E., Casstevens T., Chen C., DeClerck G., Derwent P., et al. (2011) Gramene database in 2010: updates and extensions. Nucleic Acids Res. 39: D1085–D1094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng H.Q., Chiang-Hsieh Y.F., Chien C.H., Hsu B.K.J., Liu T.L., Chen C.N.N., et al. (2014) AlgaePath: comprehensive analysis of metabolic pathways using transcript abundance data from next-generation sequencing in green algae. BMC Genomics 15: 196. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.