Fig. 3.
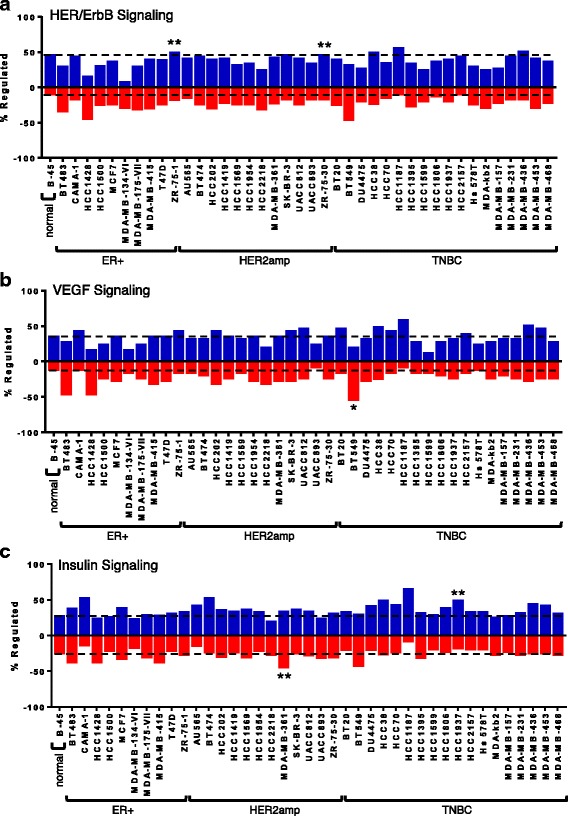
Kinome results of (a) HER/ErbB, (b) vascular endothelial growth factor (VEGF) and (c) insulin signaling pathways. For each cancer cell line, pathway overrepresentation analysis was performed using InnateDB relative to a control representing the averaged signaling profile of four non-cancer cell lines (184B5, MCF10A, MCF12A, and MCF10F; noted as normal, B-45). The percentage of peptides with increased or decreased phosphorylation is calculated relative to the total number of peptides on the array that are associated with the signaling pathway under consideration. Consideration is limited to peptides with consistent patterns of phosphorylation across the nine technical replicates (P < 0.05) and changes in the extent of phosphorylation are determined as differential phosphorylation (P < 0.05) relative to the control. The pathway overrepresentation analysis of InnateDB also provides P values for the activation or repression of the signaling pathway based on both the number of peptides that are consistently and differentially phosphorylated between the cancer cell line and the control cells and the magnitude of this phosphorylation difference. Cell lines with activation of the pathway are represented above the vertical axis (blue) or below for those with repression (red) (*P < 0.10; **P < 0.05). Dashed lines indicate the level of phosphorylation for the four averaged normal cell lines (i.e., B-45) to help identify differences for the cancer cell lines. HER2amp human epidermal growth factor receptor 2-amplified, ER estrogen receptor, TNBC triple negative breast cancer
