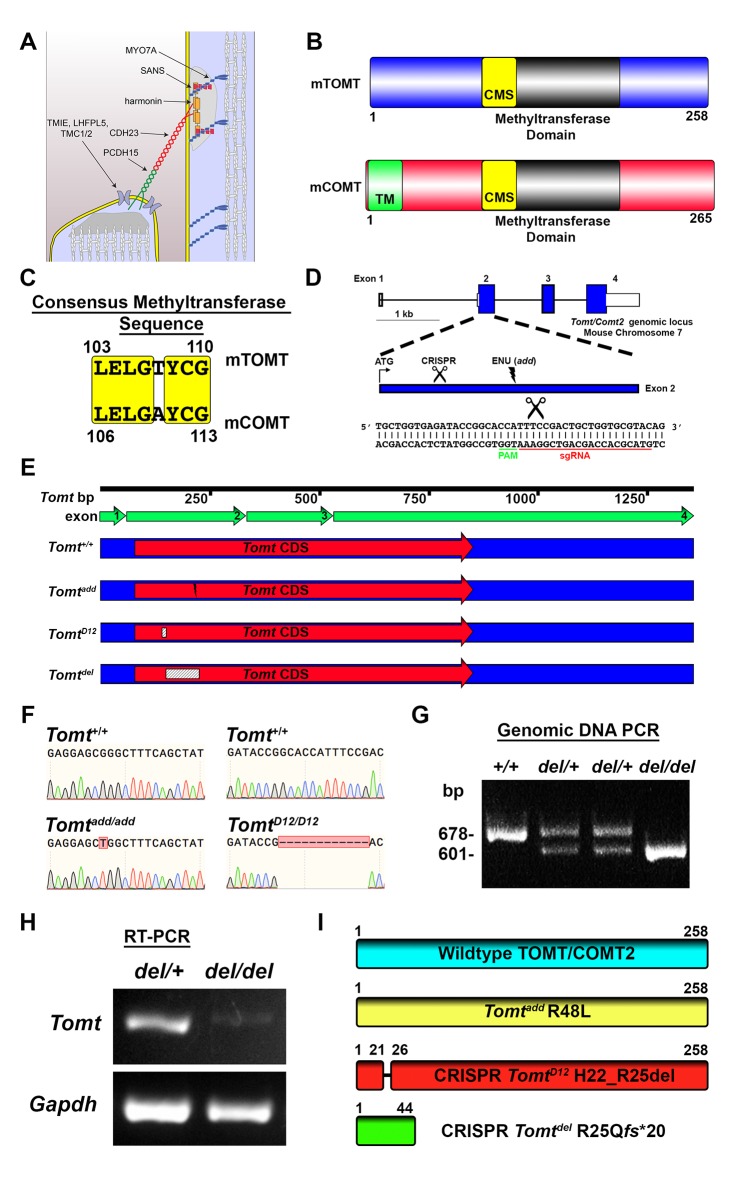
Figure 1. Generation of Tomt mutant mice.
(A) Schematic of the tip-link complex in hair cell stereocilia. (B) Schematic comparison of mouse TOMT (mTOMT, Uniprot AIY9I9) and mouse COMT (mCOMT, Uniprot O88587). Putative methyltransferase domain is indicated in black. Amino acid numbers are below. Consensus methyltransferase sequence (CMS, [Vidgren et al., 1994]) is depicted in yellow. Transmembrane domain (TM) from membrane-bound isoform of COMT is in green. (C) Conserved amino acid residues in consensus methyltransferase sequence of mTOMT and mCOMT. (D) Genomic locus of mouse Tomt (NCBI gene 791260), located on chromosome 7. Four Tomt exons are depicted as open boxes, with coding exons in blue. Introns are shown as solid black line. Exon 2, the first coding exon, is enlarged below, with relative locations indicated for ATG, clustered regularly interspaced short palindromic repeats (CRISPR) sgRNA recognition site, and add N-ethyl-N-nitrosourea (ENU) mutation site (Du et al., 2008). At bottom is Tomt exon 2 sequence showing CRISPR sgRNA and protospacer adjacent motif (PAM), and site of Cas9 cleavage (scissors). (E) Schematic of Tomt mRNA (NCBI NM_001081679.1) and location of mutations. Tomt exons are indicated with green arrows. Consensus Tomt coding sequence (CDS, NCBI CCDS40044.1) is in red. Location of add mutation is indicated with lightning bolt. Two unique deletions were identified in founder mice after CRISPR/Cas9 pronuclear injection of Tomt-specific sgRNA and Cas9. Genetic mouse lines were generated containing either a 12 base-pair (bp) deletion (TomtD12) or 77 bp deletion (Tomtdel) in Exon 2 of Tomt. Deletions are indicated as dashed-line boxes. (F) Genomic DNA sequencing results from Tomt+/+, Tomtadd/add, and TomtD12/D12 mice demonstrating mutations. (G) PCR of Tomt genomic DNA from Tomt+/+, Tomtdel/+, and Tomtdel/del demonstrating 77 bp deletion. (H) RT-PCR results for Tomt and Gapdh from inner ear tissue from Tomtdel/+ (+/−) and Tomtdel/del (−/−). (I) Predicted protein structure of wild-type and mutant TOMT. From top: wild-type TOMT; add mutation (R48L); CRISPR 12 bp in-frame deletion (TomtD12, H22_R25 del) leading to loss of four animo acids; and CRISPR 77 bp deletion (Tomtdel, R25Qfs*20) leading to a frame-shifted amino acid sequence, premature stop codon and truncated protein.