Fig. 2.
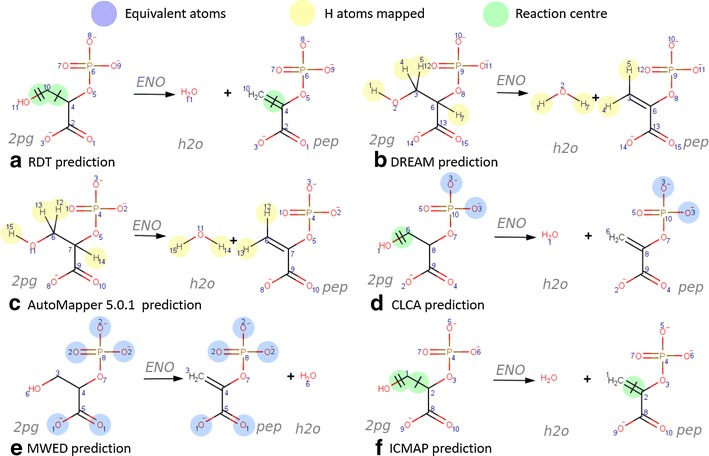
Atom mapping predictions for the enolase reaction. All six compared algorithms returned an accurate atom mapping but included different types of additional information. CLCA and MWED identify equivalent atoms in the reactants (blue). DREAM and AutoMapper map hydrogen atoms (yellow). RDT, CLCA, ICMAP and MWED all identify reaction centres (green). Unlike the other three algorithms, MWED does not identify reaction centres by adding information to the bonds that break and form. Instead, it assigns different colours to the molecular substructures (moieties) that break apart or bind together. The atom mapped reactions are visualised with MarvinView (ChemAxon, Budapest, Hungary), which accepts the RXN and SMILES formats as input
