Abstract
The florfenicol-chloramphenicol exporter gene fexA is part of the novel transposon Tn558 from Staphylococcus lentus. Similarities between Tn558 and Tn554 from Staphylococcus aureus included the arrangement of the transposase genes tnpA to -C and an att554-like target sequence. Circular forms of Tn558 were detected and suggest the functional activity of this transposon.
Recently the first staphylococcal florfenicol-chloramphenicol efflux gene, fexA, was detected on plasmid pSCFS2 of Staphylococcus lentus (4). To investigate whether fexA is part of a transposable element, the fexA flanking sequences in pSCFS2 were analyzed with regard to similarities to other staphylococcal transposons.
Plasmid pSCFS2 was digested with BglII, and the six resulting fragments of sizes between 1.2 and 14.0 kb were cloned separately into the BamHI-digested vector pBluescript II SK(+) (Stratagene, Amsterdam, The Netherlands). Confirmation of the fexA gene on the 7.1-kb BglII fragment was done by PCR and by hybridization experiments (data not shown). The PCR primers fexA-fw (5′-GTACTTGTAGGTGCAATTACGGCTGA-3′) and fexA-rev (5′-CGCATCTGAGTAGGACATAGCGTC-3′) (amplicon size, 1,272 bp; annealing temperature, 57°C) were used along with Pwo polymerase (Peqlab, Erlangen, Germany). To determine a sufficiently long sequence up- and downstream of fexA, sequence analysis included parts of the 7.1-kb BglII fragment and the adjacent 14.0-kb BglII fragment. The nucleotide sequence of the fexA flanking regions was determined by primer walking on both strands starting from the terminal parts of the fexA gene (MWG-Biotech, Ebersberg, Germany).
Analysis of a 7,718-bp region revealed the presence of a transposon-like element of 6,644 bp, designated Tn558. This element consisted of five reading frames of more than 120 amino acids (aa), which accounted for 78.5% of the Tn558 sequence (Fig. 1a). Three of these reading frames exhibited similarities to the genes tnpA, tnpB, and tnpC, whose products are involved in the transposition of the Staphylococcus aureus transposons Tn554 (1, 5-8) and Tn5406 (3) (Fig. 1a). It is noteworthy that despite the same arrangement and the similar sizes, the tnpA, tnpB, and tnpC genes of the three transposons Tn554, Tn5406, and Tn558 differed considerably in their nucleotide sequences and in the deduced amino acid sequences of their gene products (Table 1). The remaining two open reading frames of Tn558 were orientated in the opposite direction and code for the 475-aa florfenicol-chloramphenicol exporter protein FexA and a putative oxidoreductase of 138 aa, respectively (Fig. 1a). The deduced amino acid sequence of the latter open reading frame exhibited 45 to 47% identity and 76% similarity to the NAD(P)H oxidoreductases of Fusobacterium nucleatum subsp. nucleatum (NP_604130) or Fusobacterium nucleatum subsp. vincentii (ZP_00143945).
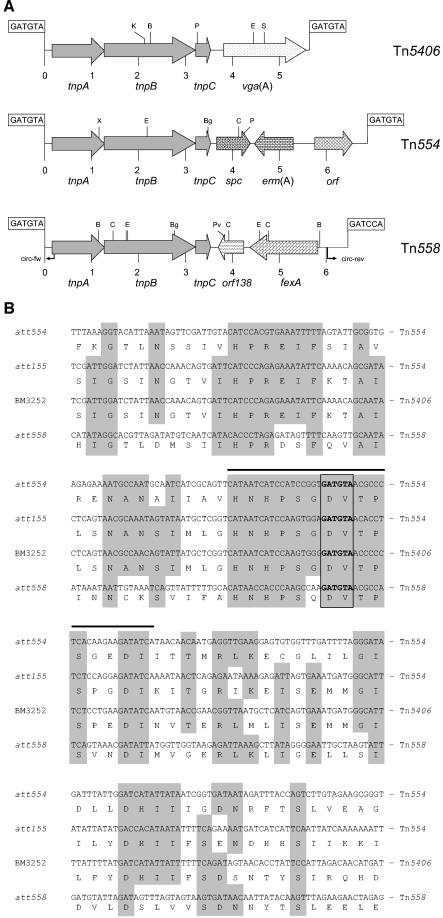
FIG.1.
(A) Organization of the S. lentus transposon Tn558 in comparison to the structurally related transposons Tn554 (X03216) and Tn5406 (AF186237). A distance scale in kilobases is given below each map. The position and orientation of the genes coding for transposition functions (tnpA, tnpB, and tnpC), antimicrobial resistance [vga(A), streptogramin A resistance; erm(A), resistance to macrolides, lincosamides, and streptogramin B antibiotics; spc, spectinomycin resistance; fexA, resistance to florfenicol and chloramphenicol], or unknown functions (orf, orf138) are indicated by arrows with the direction of transcription shown by the arrowhead. The restriction endonuclease cleavage sites are abbreviated as follows: B, BclI; Bg, BglII; C, ClaI; E, EcoRI; K, KpnI; P, PstI; Pv, PvuII; X, XhoI. The positions of primers used for the detection of circular Tn554 forms are labeled circ-fw and circ-rev and are indicated by arrows. The 6-bp core nucleotide sequences at the transposon junctions are shown in boxes. (B) Nucleotide and amino acid sequence alignment of the attachment sites att554 (in S. aureus N315 [3) and att155 (in S. epidermidis [13]) of Tn554, that of Tn5406 in S. aureus strain BM3252 (3), and att558 of Tn558 in plasmid pSCFS2. An attachment site identical to att554 has also been reported for Tn5406 in S. aureus strain BM3327 (3). Grey boxes indicate identical amino acids found in three or more of the aligned sequences. The hexanucleotide core sequences of the integration sites are framed. The black bar above the att554 sequence indicates the minimum sequence required for transposition into this site as determined by deletion analysis (5, 6).
TABLE 1.
Comparisons between the transposase genes tnpA, -B, and -C and the corresponding proteins of Tn554, Tn5406, and Tn558
| Transposon | Characteristic or comparison result for:
|
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
tnpA
|
tnpB
|
tnpC
|
||||||||||
| Size
|
% Identitya
|
Size
|
% Identitya
|
Size
|
% Identitya
|
|||||||
| Gene (bp) | Protein (aa) | Gene | Protein | Gene (bp) | Protein (aa) | Gene | Protein | Gene (bp) | Protein (aa) | Gene | Protein | |
| Tn554 | 1,086 | 361 | 100.0 | 100.0 | 1,893 | 630 | 100.0 | 100.0 | 378 | 125 | 100.0 | 100.0 |
| Tn5406b | 1,086 | 361 | 69.8 | 72.5 | 1,899 | 632 | 71.4 | 73.0 | 384 | 127 | 65.1 | 62.3 |
| Tn558b | 1,086 | 361 | 69.7 | 72.3 | 1,920 | 639 | 70.0 | 71.4 | 366 | 121 | 62.7 | 57.6 |
Identitiy with respect to the tnp gene and Tnp protein of Tn554.
Comparisons between Tn5406 and Tn558 revealed the following percentages of identity at the nucleotide/amino acid sequence level: for tnpA, 70.2/71.7; for tnpB, 69.5/71.2; for and tnpC, 60.9/56.4.
As previously reported for Tn554 and Tn5406 (3, 9), Tn558 also did not contain inverted repeats at its ends and failed to generate a duplication of the target sequence at the integration site. Tn558 exhibited the hexanucleotide sequence 5′-GATGTA-3′ at the left-end junction and a similar sequence, 5′-GATCCA-3′, at the right-end junction. The sequence 5′-GATGTA-3′ has previously been described as the “core” sequence of Tn554 and Tn5406 in the primary target site att554 in the S. aureus chromosome (Fig. 1b) (2, 3, 11-13). Studies on serial transposition of Tn554 into primary and secondary target sites revealed that the sequences at the junctions of Tn554 varied with respect to the target sites: with each new transposition event, the sequence originally present in the target site is found at the left end of Tn554, whereas the former left-end junction is now found at the right end and the former right-end junction is lost (5, 10). A similar process is assumed to be responsible for the altered sequence found at the right-end junction of Tn558 in plasmid pSCFS2. Analysis of the regions flanking the Tn558 insertion in plasmid pSCSF2 identified a sequence similar to that of att554. A comparison of this pSCFS2 region, designated att558, with the sequences up- and downstream of insertion sites of Tn554 and Tn5406 is shown in Fig. 1b. The att554 sites of Tn554 and Tn5406 are located within reading frames for proteins of 222 aa which show similarity to DNA repair proteins (3). The reading frame including the att558 site codes for a putative protein of 140 aa which showed 51% identity and 68% similarity to a 147-aa DNA repair protein from Listeria monocytogenes (ZP_00231288).
Since transposition of Tn554 and Tn5406 includes the formation of circular forms which precede the integration of the transposon into a new target sequence (3, 5), inverse PCR assays were conducted to detect these circular intermediates. For this, the Pwo polymerase (Peqlab) and the two primers circ-fw (5′-CGGTGCCTAATCATTCGTATGC-3′) and circ-rev (5′-CGCTTAACCGGTTCTATCACTTCA-3′) (amplicon size, 871 bp; annealing temperature, 62°C) were used; the primers positions are shown in Fig. 1a. Amplicons of the expected size were obtained in repeated experiments from several different S. aureus RN4220:pSCFS2 transformants (data not shown). The sequence of such amplicons (MWG-Biotech) consisted of 229 bp of tnpA and its upstream region including the 6-bp core sequence (5′-GATGTA-3′) at the left end of Tn558, whereas the remaining 642 bp of the amplicon represented the right end of Tn558 up to but not including the sequence 5′-GATCCA-3′. Evidence of the presence of circular Tn558 forms suggested the functional activity of this transposon in staphylococci (3, 5). Based on the aforementioned transposition model, this observation also suggested that the sequence 5′-GATGTA-3′ at the left-end junction of Tn558 might be part of the att558 insertion site.
The data presented in this study showed that the 6,644-bp transposon Tn558 is a member of the Tn554 family of staphylococcal transposons. Although the members of this transposon family share the same overall structure and mode of transposition, they differ distinctly in their resistance gene regions. The macrolide-lincosamide-streptogramin B resistance gene erm(A) and the spectinomycin resistance gene spc in Tn554 (8, 14) were replaced by a variant of the streptogramin A resistance gene vga(A) in Tn5406 (3) and by the florfenicol-chloramphenicol exporter gene fexA and a putative oxidoreductase gene in Tn558 (Fig. 1a). The identification of fexA as part of a functionally active transposon is an important observation with regard to the mobility of fexA and the spread of combined resistance to florfenicol and chloramphenicol. Although the novel transposon is nonconjugative, its location on a plasmid underlines the role of plasmids as vectors for transposon-borne resistance genes in the spread of antibiotic resistance.
Nucleotide sequence accession number.
The sequence of Tn558 and its flanking regions has been deposited in the EMBL database under accession number AJ715531.
Acknowledgments
This study was supported by grants of the Deutsche Forschungsgemeinschaft (SCHW 382/6-1 and SCHW 382/6-2).
We thank Vera Nöding for excellent technical assistance.
REFERENCES
- 1.Bastos, M. C., and E. Murphy. 1988. Transposon Tn554 encodes three products required for transposition. EMBO J. 7:2935-2941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Chikramane, S. G., P. R. Matthews, W. C. Noble, P. R. Stewart, and D. T. Dubin. 1991. Tn554 inserts in methicillin-resistant Staphylococcus aureus from Australia and England: comparison with an American methicillin-resistant group. J. Gen. Microbiol. 137:1303-1311. [DOI] [PubMed] [Google Scholar]
- 3.Haroche, J., J. Allignet, and N. El Solh. 2002. Tn5406, a new staphylococcal transposon conferring resistance to streptogramin A and related compounds including dalfopristin. Antimicrob. Agents Chemother. 46:2337-2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kehrenberg, C., and S. Schwarz. 2004. fexA, a novel Staphylococcus lentus gene encoding resistance to florfenicol and chloramphenicol. Antimicrob. Agents Chemother. 48:615-618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Murphy, E. 1990. Properties of the site-specific transposable element Tn554, p. 123-135. In R. P. Novick (ed.), Molecular biology of the staphylococci. VCH Publishers, New York, N.Y.
- 6.Murphy, E. 1989. Transposable elements in gram-positive bacteria, p. 269-288. In D. E. Berg and M. M. Howe (ed.), Mobile DNA. ASM Press, Washington, D.C.
- 7.Murphy, E. 1983. Inhibition of Tn554 transposition: deletion analysis. Plasmid 10:260-269. [DOI] [PubMed] [Google Scholar]
- 8.Murphy, E., L. Huwyler, and M. C. F. Bastos. 1985. Transposon Tn554: complete nucleotide sequence and isolation of transposition-defective and antibiotic-sensitive mutants. EMBO J. 4:3357-3365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Murphy, E., and S. Löfdahl. 1984. Transposition of Tn554 does not generate a target duplication. Nature 307:292-294. [DOI] [PubMed] [Google Scholar]
- 10.Murphy, E., S. Phillips, I. Edelman, and R. P. Novick. 1981. Tn554: isolation and characterization of plasmid insertions. Plasmid 5:292-305. [DOI] [PubMed] [Google Scholar]
- 11.Murphy, E., E. Reinheimer, and L. Huwyler. 1991. Mutational analysis of att554, the target of the site-specific transposon Tn554. Plasmid 26:20-29. [DOI] [PubMed] [Google Scholar]
- 12.Phillips, S., and R. P. Novick. 1979. Tn554—a site-specific repressor-controlled transposon in Staphylococcus aureus. Nature 278:476-478. [DOI] [PubMed] [Google Scholar]
- 13.Tillotson, L. E., W. D. Jenssen, L. Moon-McDermott, and D. T. Dubin. 1989. Characterization of a novel insertion of the macrolides-lincosamides-streptogramin B resistance transposon Tn554 in methicillin-resistant Staphylococcus aureus and Staphylococcus epidermidis. Antimicrob. Agents Chemother. 33:541-550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Townsend, D. E., S. Bolton, N. Ashdown, D. I. Annear, and W. B. Grubb. 1986. Conjugative staphylococcal plasmids carrying hitch-hiking transposons similar to Tn554: intra- and interspecies dissemination of erythromycin resistance. Aust. J. Exp. Biol. Med. Sci. 64:367-379. [DOI] [PubMed] [Google Scholar]