Fig. 1.
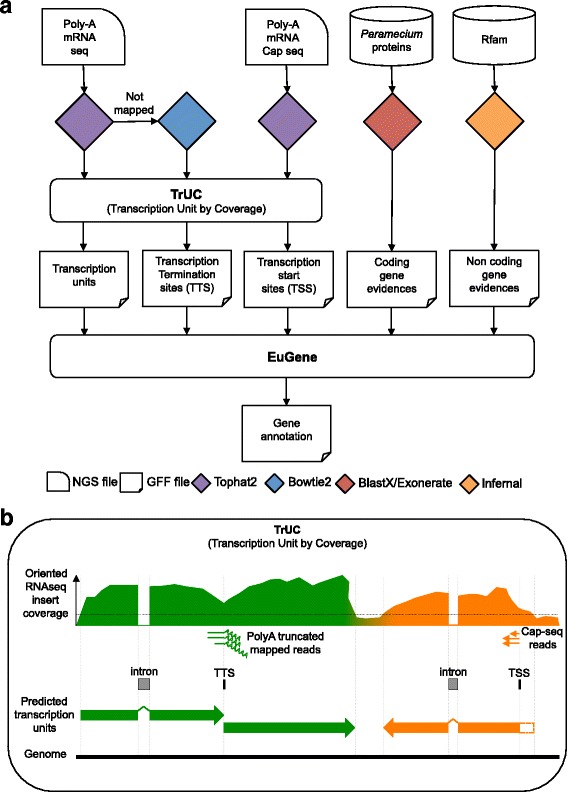
Gene annotation strategy. a Overview of the workflow. EuGene software, using a Paramecium-trained matrix, combines (i) transcription unit predictions, (ii) TSS predicted positions, (iii) TTS predicted positions, (iv) Paramecium predicted proteins mapped on the reference genome using BLASTX then Exonerate, and (v) non-coding gene predictions obtained using the Rfam database. b Schema of the TrUC pipeline. TrUC is able to predict transcription units, TSS and TTS positions. To achieve this, the software uses oriented polyA+ mRNA-Seq and Cap-Seq data. The upper part of the schema represents RNA-Seq insert coverage of the genome. A configurable threshold (horizontal dotted line) is used to determine the edges of the transcription units. The middle of the schema shows how intron, TSS and TTS positions are predicted. The transcription units predicted by combining all of the information are shown at the bottom of the schema. The TSS and the TTS are used to refine the structure of the transcription unit predictions. This can be particularly critical in a compact genome to avoid fusing adjacent transcription units. An example is shown in orange, where the TSS is used to shorten the predicted transcription unit, removing the open box. The example in green, shows how a TTS can prevent fusion of two adjacent transcription units
