Fig. 3.
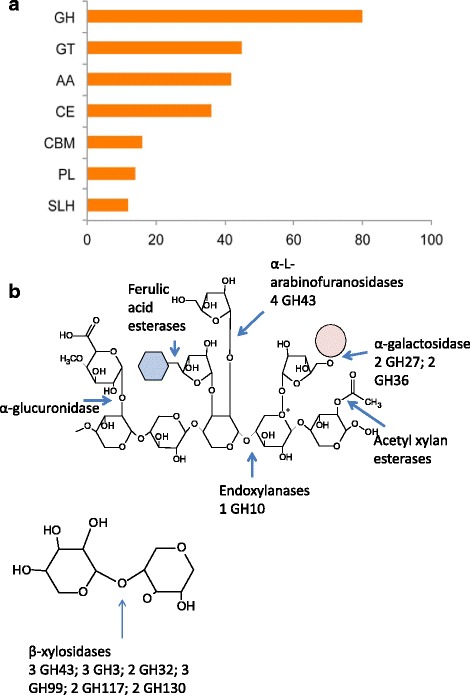
HGT genes with functional domains involved in carbohydrate metabolism as depicted from Carbohydrate-Active enZYmes Database (CAZy). a CAZy general categories among HGTs. Horizontal bars represent the number of unique sequences annotated with CAZyme domains. GH – glycoside hydrolases; GT – glycosyltransferases; AA – auxiliary activity module; CE – carbohydrate esterases; CBM – carbohydrate binding module; PL – polysaccharide lyases; SLH – S-layer homology module. b Hemicellulose degradation. The numbers represent the number of identified HGT enzymes. Pink circle represents D-Galactose; Blue hexagon represents Furelic acid
