Fig. 2.
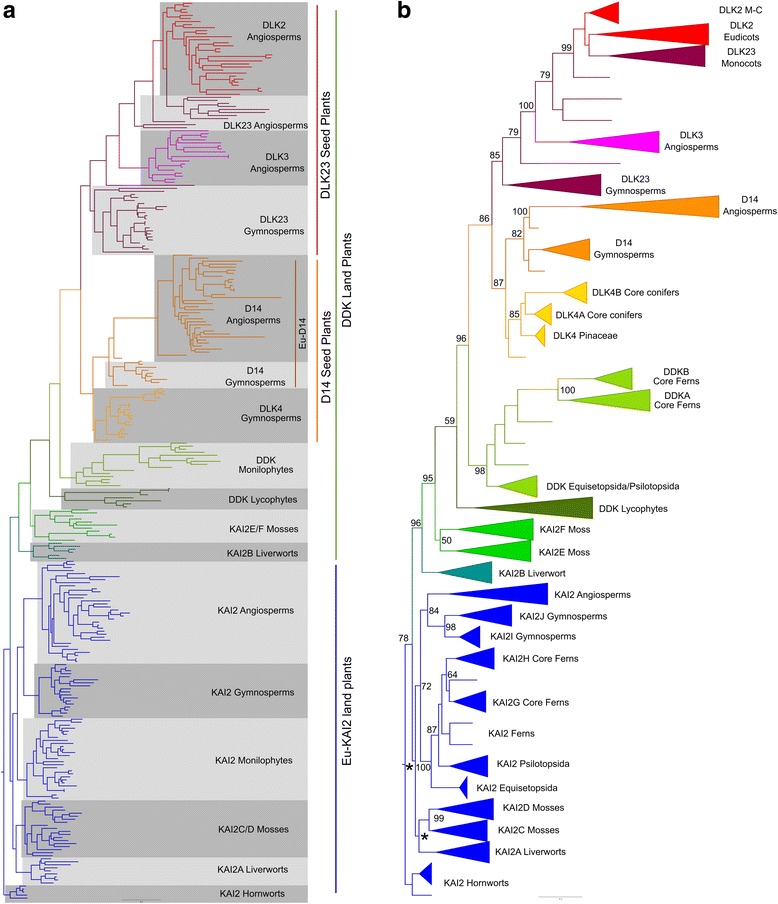
The eu-KAI2 and DDK super-clades diverged early in land plant evolution. Nucleotide-level phylogenetic analysis implemented in GARLI on the D14/KAI2 family, minus charophyte and lycophyte KAI2 sequences (296 sequences). Trees were rooted with hornwort KAI2 sequences by comparison with Fig. 1. This analysis was performed using the full-length dataset (780 characters). a Phylogram showing the ‘most likely’ tree from GARLI analysis, labelled to show the high-order relationships between the major clades (as described in Table 1). b Cladogram depicting the phylogenetic tree from (a) in simplified form. Major clades and sub-clades (as listed in Table 1) are collapsed. Numbers associated with internal branches denote maximum likelihood bootstrap support (percent support); values below 50 are indicated by *. M-C magnoliids/chloranthales
