Figure 1.
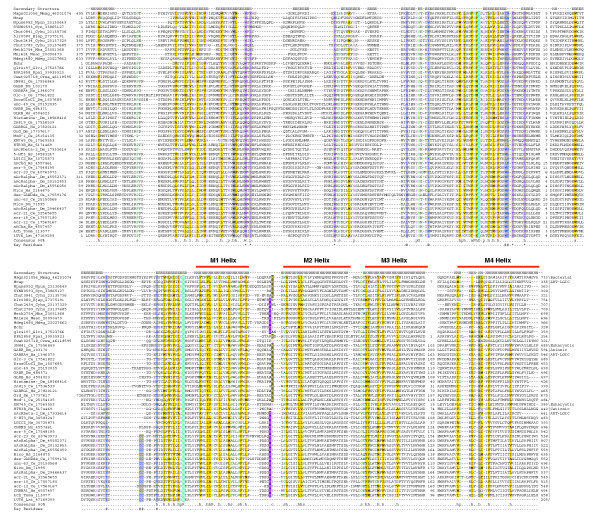
A multiple alignment of the ART-LGIC/Cys-loop superfamily (see also Additional data file 2 for alignment; an alignment of metazoan members only may also be obtained from PFAM: PF02931 LBDs; PF02932: TM domain). Proteins are denoted by their gene names, species abbreviations and gi. The secondary-structure assignments, based on the available crystal structures of the acetylcholine receptor pore (pdb: 1OED) and Achbp protein (pdb: 1UV6), are shown above the alignment where E denotes extended or strand, and H, helix. The coloring reflects the composition of the amino acids at 90% consensus. The coloring scheme and the consensus abbreviations are as follows: h, hydrophobic (ACFILMVWY), l, aliphatic residues (ILV), and a, aromatic residues (FHWY) are shaded yellow; s, small (AGSVCDNPT) and u, tiny residues (GAS) are colored green; c, charged (DEHKR), +, basic (HKR), -, acidic (DE), p, polar (CDEHKNQRST) are colored magenta. The conservation pattern as plotted onto the three-dimensional structure of the ACHB is shown in Additional data file 1. Also shown below the alignment are the key residues described in the text. # and @ represent residues of adjacent chains (PDB id: 1UV6, chain C and chain D respectively) involved in ligand binding (shaded gray). Residues predicted to be potentially involved in the transmission of conformational change are marked by an asterisk (*) at the bottom of the column and are colored violet and shaded gray. The highly conserved positions - the acidic residue in the middle of the Cys-loop and the basic residue at the carboxyl terminus of the LBD - are shown in inverse blue shading. The arginine residue involved in ion selectivity in anionic channels is shaded green and the glutamate residue involved in ion selectivity in cationic channels is shaded purple. Species abbreviations are as follows: Bjap, Bradyrhizobium japonicum; Ce, Caenorhabditis elegans; Crwa, Crocosphaera watsonii; Cyhu, Cytophaga hutchinsonii; Dm, Drosophila melanogaster; Echr, Erwinia chrysanthemi; Glvi, Gloeobacter violaceus; Hs, Homo sapiens; Lst, Lymnaea stagnalis; Mba, Methanosarcina barkeri; Mcap, Methylococcus capsulatus; Mdeg, Microbulbifer degradans; Meac, Methanosarcina acetivorans; Mmag, Magnetospirillum magnetotacticum; Npun, Nostoc punctiforme; Rpal, Rhodopseudomonas palustris; Syn, Synechococcus sp.; Toma, Torpedo marmorata.
