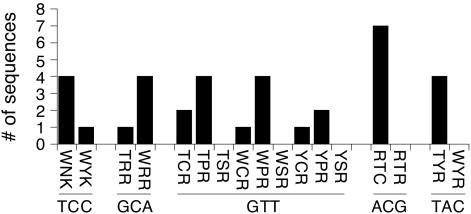
Figure 3.
Histogram showing active zinc finger proteins identified by the in vivo (cell-based) assay. ‘Mini libraries’ encoding all potential candidate proteins identified from processed zinc finger library data (Table 1) were generated by MAX randomization and their interactions in vivo with their respective target DNAs assessed by yeast one-hybrid analysis as described in Materials and Methods. For each target DNA sequence, positively staining blue colonies were picked and sequenced to establish the identity of the amino acids encoded at the positions −1, 3 and 6 of zinc finger 2. On the abscissa, vertical lettering indicates amino acid identity and horizontal lettering indicates the target DNA sequence.