Figure 1.
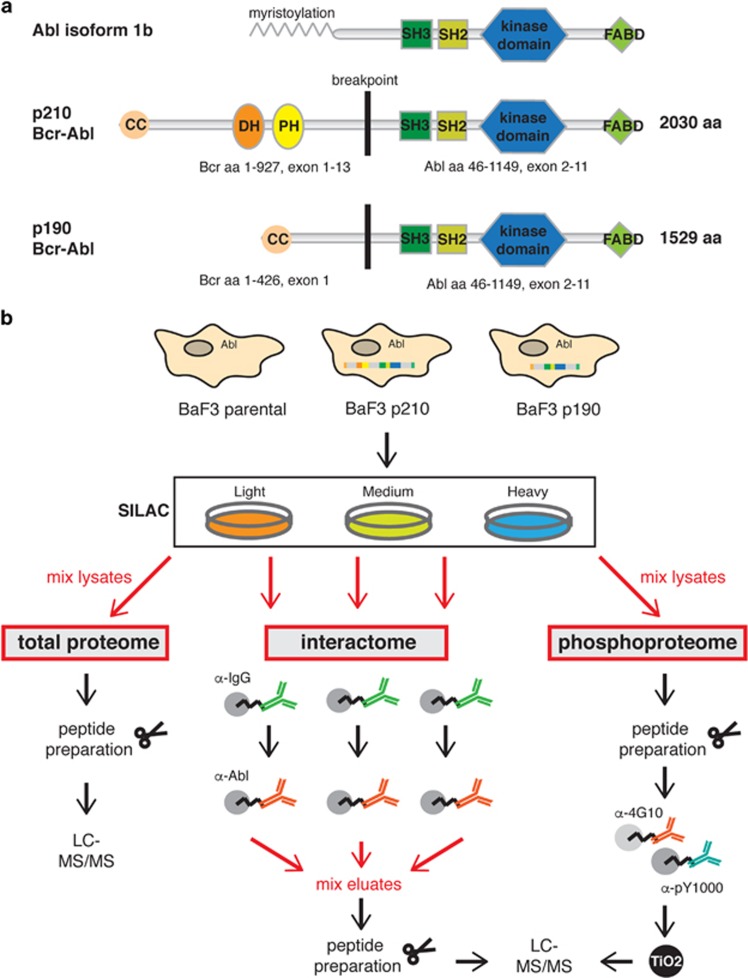
Bcr–Abl domain organization and workflow of the proteomics experiments. (a) The Abl tyrosine kinase and the two isoforms of the fusion protein Bcr–Abl, p210 and p190, are shown with their sizes and domain arrangement. The p210 isoform is 501 amino acids longer than p190 as it contains the DH–PH tandem domain. Domain abbreviations: CC, coiled-coil; DH, Dbl-homology; PH, Pleckstrin-homolgy; SH3/SH2, Src-homology 3/2; FABD, F-actin binding domain. (b) SILAC labeling was employed to allow quantitative comparison of three BaF3 cell lines (Supplementary Table S1). BaF3 parental cells express Abl endogenously. BaF3 p210 and BaF3 p190 cells express human Bcr–Abl p210 and p190. An immunoaffinity purification strategy was used to enrich for Bcr–Abl complexes for the interactome analysis and sample mixing was performed just prior to peptide preparation. For analysis of the tyrosine phosphoproteome cell lysates were mixed prior to enrichment of the pY peptides using the pY1000 and 4G10 antibodies and an additional TiO2 purification step. For both experiments, the analysis of the total proteome served for different normalization steps. LC-MS, liquid chromatography-mass spectrometry.