Fig. 3.
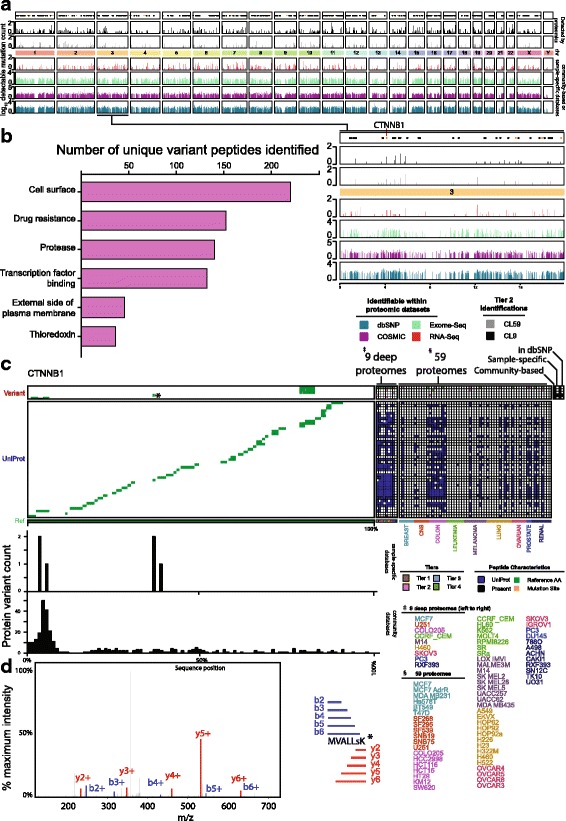
Identification of cancer-related variant peptides. a Genome coverage of potentially detectable proteogenomic peptides (6–35 amino acids) within the generated search databases (bottom). Variant proteins identified at tier 2 within 59 shallow and nine deep proteomes have been summarized in black and gray, respectively (top). Black dots correspond to the locations of COSMIC cancer census genes and orange dots indicate those detected at tier 2. b Variants identified were assessed by the drug gene interaction [43] database to identify variants that might potentially be targetable or affect related pathways. Counts relate to the number of variant peptides identified in each category for tier 2 peptides. Only categories significantly enriched at p < 0.01 are depicted. c Variant peptides detected for CTTNB1. Mutation locations have been depicted in orange. Identification of reference peptides for the same protein are shown in blue, with an alignment describing the peptides detected. Bar plots illustrate the variants that were present in genomics for this gene (top) and all mutations present in community-based databases (bottom). d A tier 2 peptide identified for CTTNB1 showing clear coverage of y and b ions
