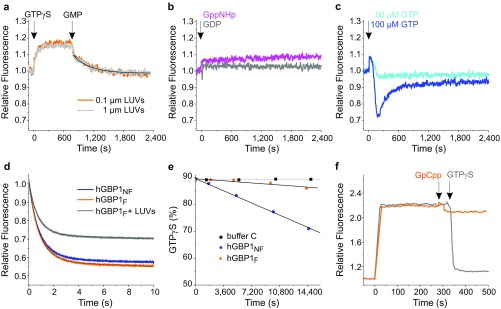
Fig. S3.
Liposome binding of hGBP1 induced by GTP analogs and their biochemical characterization. (A–C) Binding of AF-Q557CF to LUVs labeled with Rhodamine lipids. At time t = 0 s, indicated nucleotides (arrows) were added into a mixture of 0.5 mg/mL of LUVs and 2.5 µM of protein and an increase of fluorescence intensity reported liposome binding. (A) Addition of 200 µM GTPγS to LUVs of various sizes, 0.1 µm (orange) compared with 1 µm (gray). After ∼800 s, GTPγS was displaced with 5 mM of GMP, leading to dissociation of the protein from LUVs. Regardless of the LUV sizes, single exponential fits (black lines) yielded similar dissociation rate constants (koff = 0.0028 s−1 orange, koff = 0.0030 s−1 gray). (B) Addition of either 400 µM GppNHp (magenta) or GDP (gray) to 0.1 µm LUVs. (C) Addition of various concentrations of GTP to 0.1 µm LUVs. The fluorescence traces of GTP-dependent liposome binding were interfered with GTP-dependent polymerization events as demonstrated by different concentrations of the substrate (cyan, 50 µM GTP; blue, 100 µM GTP). (D) Farnesyl- and membrane-dependent dissociation kinetics of hGBP1 and GTP were investigated with hGBP1NF (blue) and hGBP1F either in the absence (orange) or presence of 2 mg/mL LUVs (gray) using the MANT-labeled GTP analog GTPγS (mGTPγS) (19). The preformed complex of 1 µM protein and 0.5 µM mGTPγS was rapidly mixed with 500 µM of nonlabeled GMP to trigger quasi-irreversible displacement of the GTP analog. The decrease of MANT fluorescence (λex = 366 nm, emission detected through 420-nm cutoff filter) reporting dissociation of mGTPγS was fitted with a single exponential equation (black lines) yielding similar dissociation rate constants under all tested conditions (koff = 1.18 s−1 blue, koff = 1.21 s−1 orange, and koff = 1.34 s−1 gray). (E) GTPγS hydrolysis catalyzed by hGBP1NF or hGBP1F was investigated with 500 µM GTPγS and 10 µM of protein. Kinetics of GTPγS turnover fitted with linear regression (solid lines) yielded specific activities of 0.037 min−1 (hGBP1NF) and 0.0058 min−1 (hGBP1F). In the absence of protein (buffer C only), the GTPγS concentration remained stable over time (dotted line). (F) Binding of the nonhydrolyzable GTP analog GpCpp to hGBP1F as tested by its ability to compete with mGTPγS used as fluorescent reporter of nucleotide binding (λex = 366 nm, λem = 435 nm). At t = 0 s, addition of 0.5 µM hGBP1F to 0.25 µM mGTPγS in the cuvette led to a fast and more than twofold increase of the fluorescence signal, indicating protein/nucleotide complex formation. Addition of 250 µM nonfluorescent GpCpp in contrast to GTPγS (arrows) did not suffice to displace the preformed protein/nucleotide complex, suggesting that hGBP1F does not bind GpCpp.