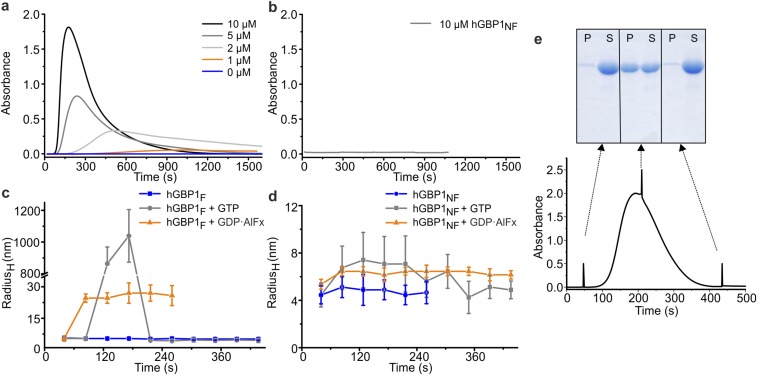
Fig. S5.
Nucleotide-dependent polymerization of hGBP1F. (A and B) Absorbance of the sample with different concentrations of hGBP1F (A) and hGBP1NF (B) after addition of 1 mM GTP (t = 0 s). (C and D) Analysis of polymer formation of 20 µM hGBP1F (C) and hGBP1NF (D) by dynamic light scattering. The particle size, given as the hydrodynamic radius (RH) in nanometers without nucleotide (blue) and in the presence of 1 mM GTP (gray) or GDP·AlFx (orange) (300 µM AlCl3, 10 mM NaF, and 200 µM GDP) was monitored over 250–450 s at 25 °C. Farnesylated hGBP1 transiently forms very large polymers during GTP hydrolysis with a radius around 1,000 nm. In contrast, the addition of GDP·AlFx leads to an irreversible formation of polymers with an average RH of 25 nm. Nucleotide-free hGBP1NF shows a RH of 5 nm. Complexation with GDP·AlFx leads to a slight irreversible increase to 6.5 nm and addition of GTP to a transient increase to a maximum value of 7.2 nm. (E) Reversibility of GTP-induced hGBP1F polymers investigated by sedimentation. During a typical turbidity experiment (see A, 10 µM hGBP1F and 1 mM GTP added at t = 0 s), samples were taken at indicated time points (arrows), spun down, and the fraction of protein in pellet (P) and supernatant (S), respectively, was analyzed by SDS/PAGE (100-µL samples were taken, centrifuged at 30,000 × g, 4 °C and for 1 min. After removal of the supernatant, the pellet was resuspended in the same volume of buffer, and corresponding fractions of protein in pellet (P) and supernatant (S) were analyzed by SDS/PAGE. Considering that the proportion of protein in the pellet represents protein complexes large enough to sediment, e.g., polymers, SDS/PAGE analysis reflected the same observation as already monitored by time-trace of absorbance: The GTP-induced increase and decrease of the absorbance considered as reversible assembly and disassembly of hGBP1F polymers was accompanied by increase and decrease of protein share in the pellet.