Figure 2.
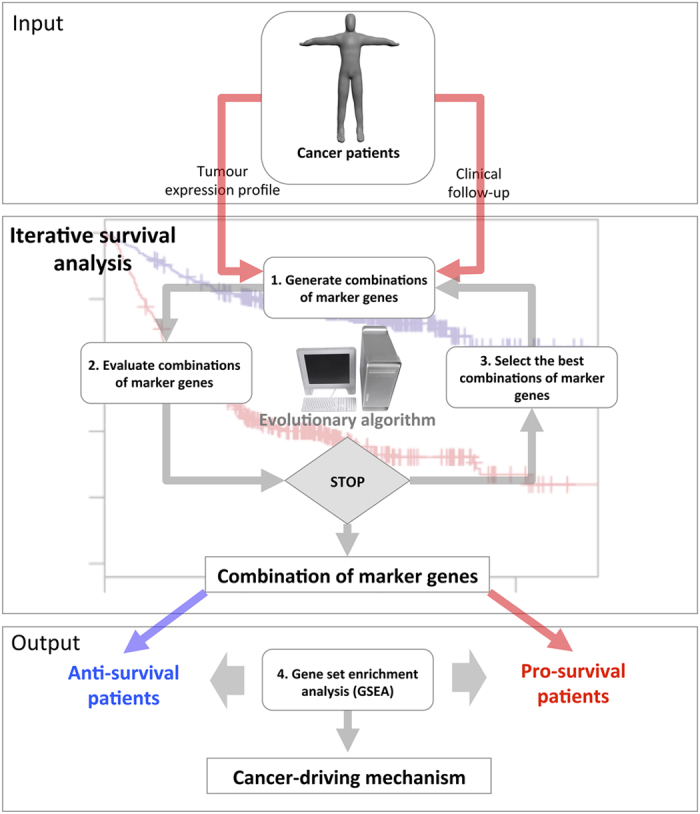
SURCOMED flow chart. SURCOMED takes as input tumor transcriptomics data and the clinical information from the corresponding patients, in particular, the survival time. The output consists of biological processes, molecular mechanisms or pathways up- or downregulated between groups of patients with long and short survival time. These groups of patients are defined by sets of marker genes identified by iterative survival analysis using an evolutionary algorithm. The iterative survival analysis can be described in 3 steps: (1) Generate combinations of marker genes. At the first iteration, the combinations are totally random; in posterior iterations, the generation of new combinations is based on the probability distribution of survival marker genes within the best combinations in the previous iteration; (2) Evaluate combinations of marker genes. This evaluation is based on the difference between the restricted mean survival time between the pro- and anti-survival groups; and (3) Select the best combinations of marker genes. Once the iterative survival analysis finishes, the resulting optimized combinations of marker genes are used to split the population of patients into pro- and anti-survival groups. A gene set enrichment analysis (GSEA) is subsequently applied in order to identify molecular mechanisms, biological processes or pathways for which their constituent genes exhibit concordant differences between pro- and anti-survival groups. This allows the identification of deregulated mechanisms between pro- and anti-survival patients.
