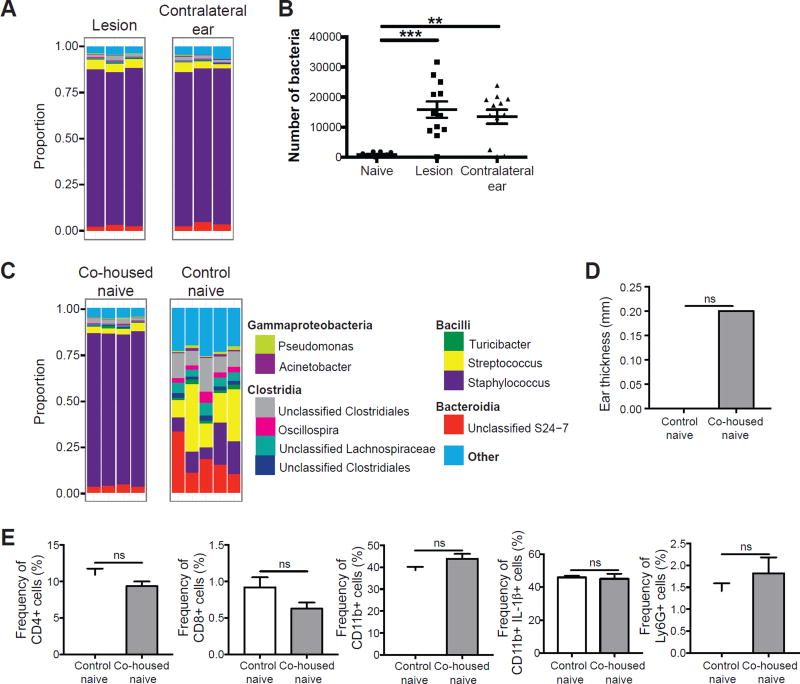
Figure 6. L. major induced dysbiosis is transmissible to uninfected skin.
(A) C57BL/6 mice were intradermally infected with L. major and swabs were collected from the infected and contralateral ears at 6 weeks post-infected for 16S rRNA gene analysis. Stacked bar charts represent the proportion of the top 10 taxa present in each sample. Data are representative of three independent experiments (n = 1 swab of each ear from 15 mice). (B) Swabs from naïve or L. major infected C57BL/6 mice were cultured on mannitol salt agar plates and CFUs were counted to determine bacteria burden. Data are representative of 1 experiment (For naïve group, n = 1 swab from the ear of 10 mice; for infected and contralateral ears, n = 1 swab of each ear from 12 mice). (C) Naïve C57BL/6 mice were co-housed with L. major infected mice for 6 weeks, while control naïve mice were housed separately. Swabs were collected from co-housed naïve and control naïve mice. Stacked bar charts represent the proportion of taxa present in each sample. Data are representative of two independent experiments (For infected group, n = 1 swab of each ear from 15 mice; for co-housed naïve, n = 1 swab of one ear from 10 mice; for control naïve, n = 1 swab of one ear from 5 mice). (D) Bar graphs depict ear thickness of control and co-housed naïve mice. (E) Cells were isolated from the ears of co-housed naïve mice and control naïve mice to assess for CD4+, CD8+, and CD11b+, IL-1β+, and Ly6G+ cells by flow cytometry. Data are representative of one experiment (Co-housed naïve, n = 1 ear tissue each from 4 mice; control naïve, n = 1 ear tissue each from 5 mice). ns = not significant; **, p < 0.01; ***, p < 0.001. See also Figure S5.