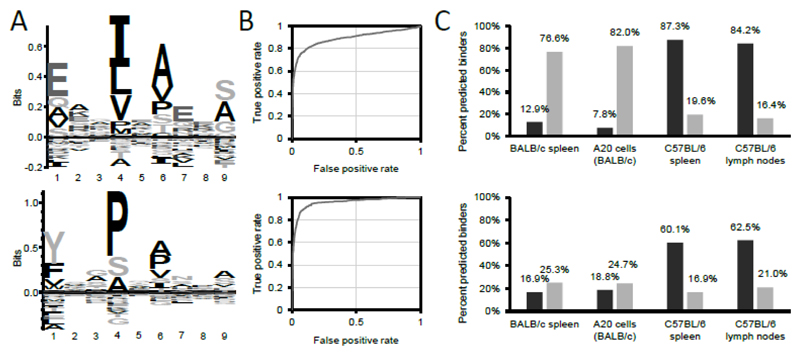
Figure 3. Matrix-based prediction of the binding of peptide sequences to mouse MHCII alleles I-Ab and I-Ad.
(A) MHC-specific motif of I-Ad (upper panel) and I-Ab (lower panel). For the creation of the motifs, I-Ad and I-Ab complexes were purified from BALB/c or C57BL/6 spleens, respectively, and eluted peptides analyzed by mass spectrometry. Peptide sequences identified with 1% FDR were clustered with GibbsCluster-1.1 server (11) and resulting I-Ad and I-Ab-specific motifs visualized with Seq2logo 2.0 (12). Seq2logo also provides positional-specific scoring matrices (PSSM, see Supplemental Material Figure 1A-B for the PSSMs) which are the basis to calculate scores. (B) ROC curves presenting the performance of the matrix-based scoring to discriminate between a true positive set (i.e. peptide sequences eluted from I-Ad (upper panel), or I-Ab (lower panel)) and a false positive set (i.e. all 15mer sequences derived from the murine reference proteome). The point at which the distance to the diagonal (i.e. the random distribution) is largest was defined as the score cut-off predicting binding to respective MHCII alleles. According to this definition, the minimum scores to predict binding were 8.0 for I-Ad, and 8.5 for I-Ab. (C) Evaluation of the matrix-based scoring (upper panel) and NetMHCIIpan 3.1 (lower panel) (15) to predict binding of peptide sequences eluted from MHCII complexes to their cognate MHCII alleles. Peptide sequences eluted from BALB/c or C57BL/6 spleens represent the training datasets used to compute the PSSM and the minimum score to predict binding. Peptide sequences from MHCII complexes of A20 cells (a B cell lymphoma cell line derived from BALB/c mice) and C57BL/6 lymph nodes represent validation data. Binding was predicted either interrogating the peptide sequences for binding to I-Ad (light gray), or to I-Ab (dark gray), with the respective PSSMs, or by selecting the respective allele on the NetMHCIIpan 3.1 server website. Peptide sequences identified from A20 cells, BALB/c and C57BL/6 spleens were taken from (7).