Fig. 3.
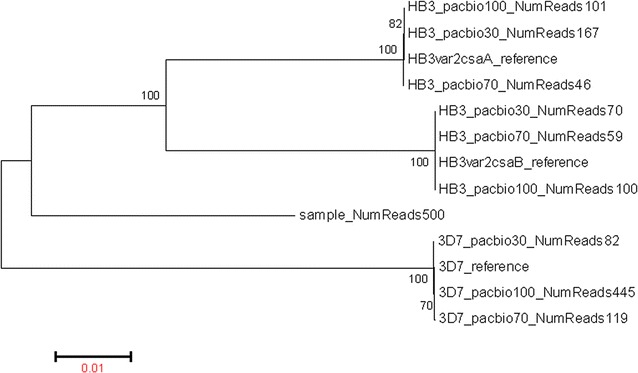
A maximum likelihood phylogenetic tree of reference and PacBio-reconstructed var2csa allelic sequences. Each sequence is named using the strain’s name (e.g. 3D7) followed by the sequencing platform (PacBio), with the proportion of the DNA composed of the specific strain (30, 70, 100%) and the number of reads used to form the consensus sequences (NumReadsxx). A phylogenetic tree of the reconstructed alleles was built using maximum likelihood, as implemented in MEGA v.6 with default parameters. The analysis showed PacBio-generated sequences to cluster with reference alleles as expected, and detected minor alleles. 3D7_PacBio denotes PacBio-sequenced 3D7, HB3_PacBio PacBio-sequenced HB3, 3D7_70 a 3D7 sequence resolved from a mixture of 70% 3D7 and 30% HB3, and HB3_70 an HB3 allele from a mixture of 30% 3D7 and 70% HB3. Of note, HB3 has two alleles of var2csa (A, B); therefore, a mixture of 3D7 and HB3 are expected to have 3 alleles. The second allele of HB3, representing 15% of the mixture in theory, was not detected in the 70% 3D7 and 30% HB3 mixture. Bootstrap support values are indicated. The scale reflects sequence divergence per base pair. 3D7_ref, HB3var2csaA, and HB3var2csaB are the allelic sequences obtained from the VarDom database
