Figure 3. IGFBP2 polypeptide overabundance in Imp2−/− MEFs is due to increased gene transcription whereas Grb14 polypeptide overabundance is due to markedly slower polypeptide degradation.
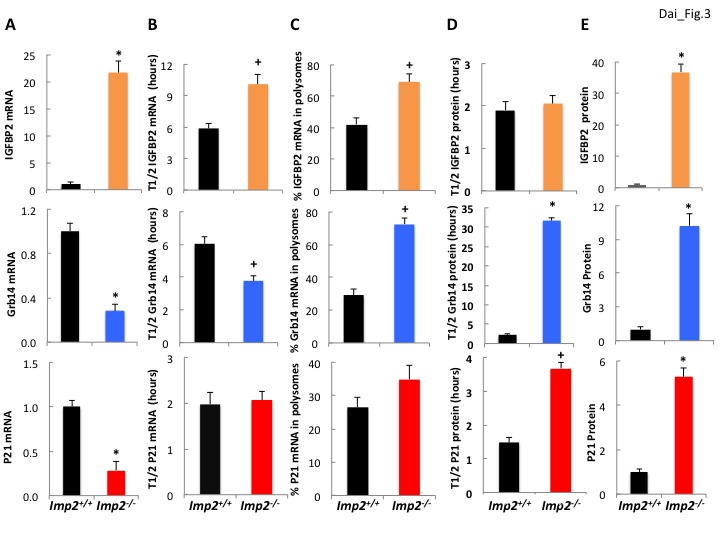
(A) Igfbp2, Grb14 and P21 mRNA abundance in Imp2+/+ and Imp2−/− MEFs. RNA was extracted from Imp2+/+ and Imp2−/− MEFs and the abundance of Grb14, Igfbp2, P21, Gapdh and Actb mRNA was determined by QPCR. The difference in Ct value between the Imp2+/+ and Imp2−/− MEFs was converted to a numerical fraction; the value for the abundance of the Imp2+/+ MEFs was set to 1.0 (black bars) and the relative abundance of the Imp2−/− MEFs is shown in colored bars. Values determined by genome-wide RNAseq are found in Supplementary file 2. (B) The turnover of Igfbp2, Grb14 and P21 mRNAs in Imp2+/+ and Imp2−/− MEFs. Actinomycin 1 uM was added to rapidly growing Imp2+/+ (black) and Imp2−/− (colored) MEFs; RNA was extracted at 0,2,4,6,8,10, 12 hr thereafter and the abundance of Grb14, Igfbp2, P21, Gapdh and Actb was determined by QPCR. The rate of decline was determined by least squares and the time representing a 50% decrease of the initial value is shown. (C) The percentage of Igfbp2, Grb14 and P21 mRNAs residing in polysomes in Imp2+/+ and Imp2−/− MEFs. Total RNA and post-mitochondrial extracts were prepared from equal numbers of rapidly growing Imp2+/+ (black) and Imp2−/− (colored) MEFs; the post-mitochondrial extracts were subjected to sucrose density gradient centrifugation. Total RNA and RNA extracted from the pooled polysomal fractions was quantified by QPCR and the ratio of polysomal/total RNA X 100 was determined for Grb14, Igfbp2 and P21. (D) The turnover of the IGFBP2, Grb14 and P21 polypeptides in Imp2+/+ and Imp2−/− MEFs. Cycloheximide 250 μM, was added to rapidly growing Imp2+/+ (black) and Imp2−/− (colored) MEFs; extracts prepared at t = 0 and every hour for 12 hr were subjected to SDS-PAGE and immunoblot for Grb14, IGFBP2 and P21. Relative abundance was determined by densitometry; the rate of decline was calculated as in 3B. (E) The abundance of the IGFBP2, Grb14 and P21 polypeptides in Imp2+/+ and Imp2−/− MEFs. Extracts prepared from rapidly growing Imp2+/+ and Imp2−/− MEFs were subjected to SDS-PAGE and the relative abundance of Grb14, IGFBP2 and P21 was determined by immunoblot; the values for Imp2+/+ were set to 1.0 (black) and the relative value for Imp2−/− is shown in the colored bars. Values determined by MS are found in Supplementary file 2. For all sections, +p<0.05, *=p < 0.01 vs Imp2+/+.
