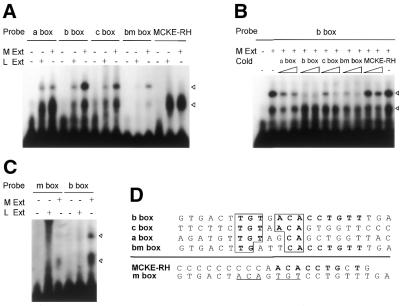
Figure 6.
A novel muscle-specific factor is responsible for recognition of TGTnACA sequences in all three 5′-distal sub-elements of the rMME of rm-hsc71m. (A) Electrophoretic mobility shift analysis showing binding specificity of a novel muscle-specific factor to rm-hsc71mE-related sequences in each sub-region of the rMME of rm-hsc71m. The rm-hsc71mE-related sequences in each region (a box, b box and c box, respectively; see panel D for each sequence) were identified by a computer search. The bm box is derived from the b box with three base changes. The MCKE-RH is the right half E-box in the MCKE (see Table 1). Each box was used as a probe in a gel shift analysis containing Rivulus muscle or liver S150 extracts. Specific and non-specific bands are marked by filled and unfilled arrowheads, respectively. (B) Competitive electrophoretic gel mobility shift analysis showing the relative binding affinity of the muscle-specific factor to each box. Increasing amounts (0, 10× and 50×) of a molar excess of cold competitor were added to the reaction containing the b box probe and Rivulus muscle S150 extracts. Specific and non-specific bands are marked by filled and unfilled arrowheads, respectively. (C) Electrophoretic mobility shift analysis showing that the central TGTnACA sequences are critical for muscle-specific factor binding. A double-stranded mutant oligonucleotide (m box; see panel D) in which all of the central conserved sequences of b box were altered (TGTGACA→ACAGTGT) was used as probe, along with parent b box probe, containing Rivulus muscle and liver S150 extracts. (D) Comparison of sequences recognized by a muscle-specific factor (only a top strand of sequences is shown: a, b and c boxes corresponded to sequences from –2627 to –2606, from –2296 to –2275 and from –2009 to –1988, respectively). Consensus binding bases are indicated as a boldface letter in a box. Boldface letters indicate homologous sequences in the MCKE-RH and the b box, and the underlined sequences in the m box indicate mutagenized bases from the b box.