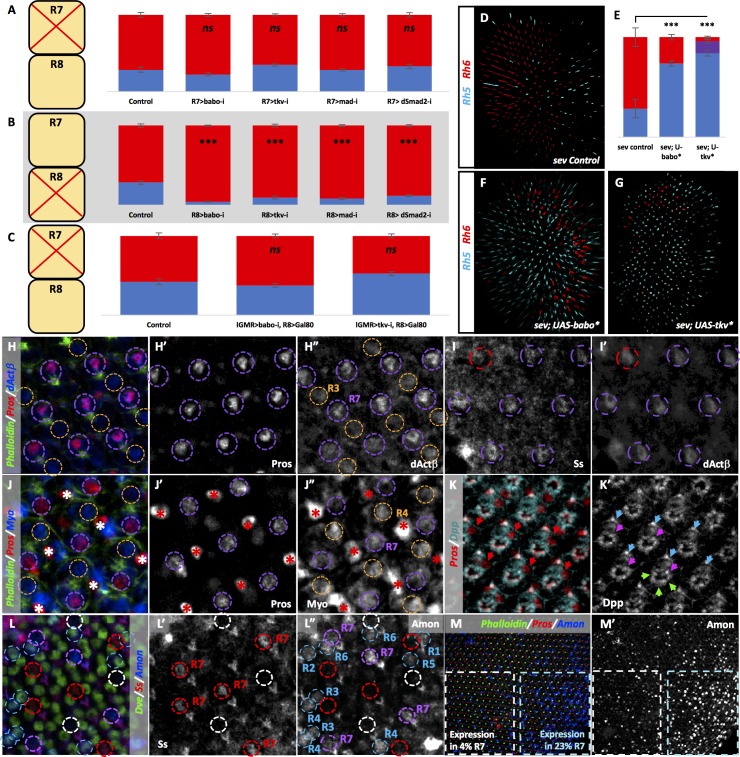
Figure 5. TGFβ ligands are processed specifically in pale R7 and activate their receptors and subsequent R-SMADs in R8.
Graph Legend: Red is the percentage of R8 photoreceptors that expressed Rh6, Blue is the percentage that expressed Rh5 and Purple is Co-expression of Rh5 +Rh6. Error bars report standard error of the mean. Paired t-tests were performed against percentages of Rh5 expression and significance is denoted where applicable (***=p ≤ 0.01, ns = not significant). Percentage values for each genotype are listed as (Rh5/(Rh5 +Rh6)/Rh6). Control for A is PM1.81-Gal4; Pros-Gal4 >UAS-Dcr2 (29/0/71) (n = 6). For B, the control is Sca-Gal4; Sens-Gal4 > UAS-Dcr2 (28/0/72) (n = 7). Control for C is lGMR-Gal4 > UAS-Dcr2; Sca-lexA > lexAOP-Gal80 (42/0/58) (n = 7). (A) Removing babo (PM1.81-Gal4; Pros-Gal4 > UAS-Dcr2; UAS-BaboRNAi) (23/0/77) (n = 6), tkv (PM1.81-Gal4; Pros-Gal4 > UAS-Dcr2; UAS-TkvRNAi) (35/0/65) (n = 7), mad (PM1.81-Gal4; Pros-Gal4 >UAS-Dcr2; UAS-MadRNAi) (28/0/72) (n = 5), or dSmad2 (PM1.81-Gal4; Pros-Gal4 >UAS-Dcr2; UAS-dSmad2RNAi) (33/0/67) (n = 6) specifically in R7 had no effect on Rh5/Rh6 ratios. (B) Removing babo (Sca-Gal4; Sens-Gal4 > UAS-Dcr2; UAS-BaboRNAi) (4/0/96) (n = 6), tkv (Sca-Gal4; Sens-Gal4 > UAS-Dcr2; UAS-TkvRNAi) (10/0/90) (n = 7), mad (Sca-Gal4; Sens-Gal4 > UAS-Dcr2; UAS-MadRNAi) (8/0/92) (n = 4), or dSmad2 (Sca-Gal4; Sens-Gal4 > UAS-Dcr2; UAS-dSmad2RNAi) (12/0/88) (n = 4) specifically in R8 reduced the number of pale subtypes as we saw with lGMR-Gal4 drivers. (C) Silencing Gal4 expression specifically in R8 attenuated the lGMR-Gal4 > UAS BaboRNAi- and TkvRNAi-induction of yellow R8 subtypes (lGMR-Gal4 > UAS-Dcr2; UAS-BaboRNAi; Sca-lexA > lexAOP-Gal80) (37/0/63) (n = 6) (lGMR-Gal4 > UAS-Dcr2; UAS-TkvRNAi; Sca-lexA > lexAOP-Gal80) (52/0/48) (n = 4). (D) sev mutations genetically ablate R7 cells resulting in increased amounts of default yellow R8 cells. Our sev mutant control, which carries a lGMR-Gal4 driver, expressed higher levels of pale subtypes (29/0/71) than observed in sev mutants in a WT background (4/0/96), likely due to genetic background (n = 3). (E) Quantification of D, F-G. (F) Expressing UAS-Babo* in a sev mutant (sev14; lGMR-Gal4 > UAS-Babo*) (74/0/26) (n = 5) increased the number of pale subtypes significantly. (G) Expressing UAS-Tkv* in a sev mutant (sev14; lGMR-Gal4 > UAS-Tkv*) (84/12/4) (n = 5) increased pale subtypes significantly. (H-M) Pupal retinas, 45–55 hr APF. (H) dActβ was expressed in all R7 cells (circled in purple) and less strongly in some R3 cells (circled in orange). (H’) Single-channel staining of the Prospero (Pros) transcription factor, which is expressed in all R7 cells. (H’). Single-channel dActβ protein reporter overlaped with Pros expression. (I) The Spineless (Ss) antibody marks yellow R7 cells (circled in purple). (I’) dActβ was expressed in both Ss-positive yellow R7 cells (circled in purple) and Ss-negative pale R7 cells (circled in red). (J) A Myo protein reporter was expressed in all R7 cells (circled in purple) as well as R4 cells (circled in orange). Bristle cells also labeled brightly (marked with asterisks). (J’) Single-channel Pros staining. (J’) Single-channel Myo reporter. (K) A Dpp-GFP transcriptional reporter was expressed in all photoreceptors. Pros marks R7 cells (red arrows). (K’) Single-channel Dpp reporter. Blue arrows mark R6 expression; green arrows mark expression in photoreceptors 1–5; pink arrows mark expression in R7. (L) An Amon-Gal4 transcriptional reporter was expressed in some pale R7 cells (circled in purple) but is absent in others (circled in white). It was absent from yellow R7 cells (circled in red). The reporter was also expressed in some outer photoreceptors (circled in blue). (L’) Single channel Ss staining marks yellow R7 cells (circled in red). (L’) Single-channel Amon reporter. Amon pale R7 expression – 9%, yellow R7 expression – 1%. N = 2678 ommatidia. (M) Amon expression was variable across individual pupal retina. (M’) Single channel Amon reporter expression.