Figure 1. Chromosome structures and SMC proteins during the cell cycle.
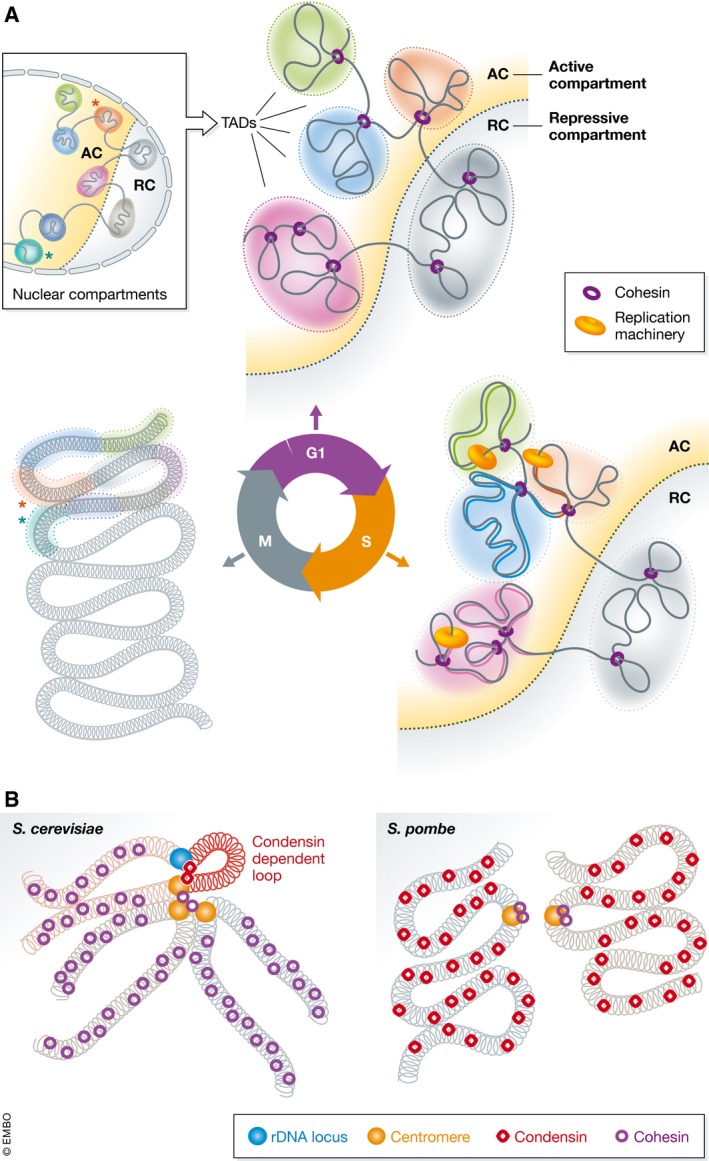
(A) Schematic representations of chromosome structure during the cell cycle. TADs on a section of a chromosome are indicated as shaded areas in active (AC) or repressive (RC) compartments, separated by the dotted line. Cohesin is shown in purple and replication machinery in orange on the DNA. In G1, TADs are insulated from one another and occupy distinct nuclear space and compartments. During S‐phase, DNA is replicated at specific times, from early to late replicating domains indicated by proportion of replicated DNA in the TAD. TAD insulation is maintained, albeit to a lesser extent, but compartmentalisation increases. Once in M‐phase, the chromatin is highly compacted with TAD structure barely identifiable and abundant very‐long‐range contacts emerge between distant TADs (e.g. compare the relationship between the orange and turquoise TADs marked with stars in the zoom‐out of G1‐phase to the M‐phase). (B) Comparison of the structure of mitotic chromosomes in yeast (for simplicity, only individual sister chromatids are shown). Saccharomyces cerevisiae chromosome arms are compacted by cohesin compared to Schizosaccharomyces pombe where condensin is required. The rDNA locus of S. cerevisiae is brought into proximity of the centromere by condensin, which is not required at other loci.
