Fig. 2.
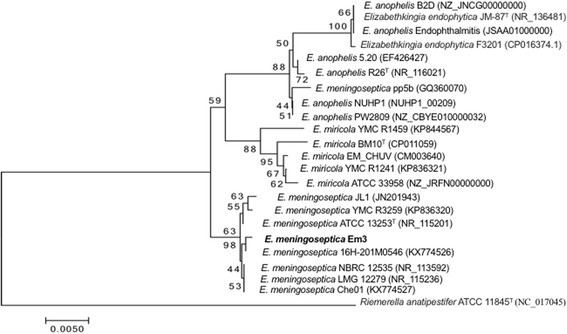
Phylogenetic tree displays the position of E. meningoseptica Em3 (shown in bold) relative to the other type strains of Elizabethkingia based on 16S rRNA. The phylogenetic tree was constructed by MEGA v. 7.0.14 using the Neighbor-Joining method [42]. The percentage of replicate trees where the associated taxa clustered together in the bootstrap test (500 replicates) is indicated next to the branches. The branch lengths are scaled to the same units as those of the evolutionary distances for inferring the phylogenetic tree. The accession numbers for 16 s rRNA sequences are listed in the parenthesis following selected bacteria: E. meningoseptica LMG 12279 (NR_115236), E. meningoseptica Che01 (KX774527), E. meningoseptica NBRC 12535 (NR_113592), E. meningoseptica ATCC 13253T (NR_115201), E. meningoseptica JL1 (JN201943), E. meningoseptica YMC R3259 (KP836320), E. meningoseptica Em3, E. meningoseptica 16H-201 M0546 (KX774526), E. miricola YMC R1459 (KP844567), E. miricola BM10T (CP011059), E. miricola EM_CHUV (CM003640), E. miricola YMC R1241 (KP836321), E. miricola ATCC 33958 (NZ_JRFN00000000), E. meningoseptica pp5b (GQ360070), E. anophelis NUHP1 (NUHP1_00209), E. anophelis PW2809 (NZ_CBYE010000032), E. anophelis 5.20 (EF426427), E. anophelis R26T (NR_116021), E. anophelis Endophthalmitis (JSAA01000000), Elizabethkingia endophytica JM-87T (NR_136481), Elizabethkingia endophytica F3201 (CP016374.1), E. anophelis B2D (NZ_JNCG00000000) and Riemerella anatipestifer ATCC 11845T (NC_017045)
