Fig. 4.
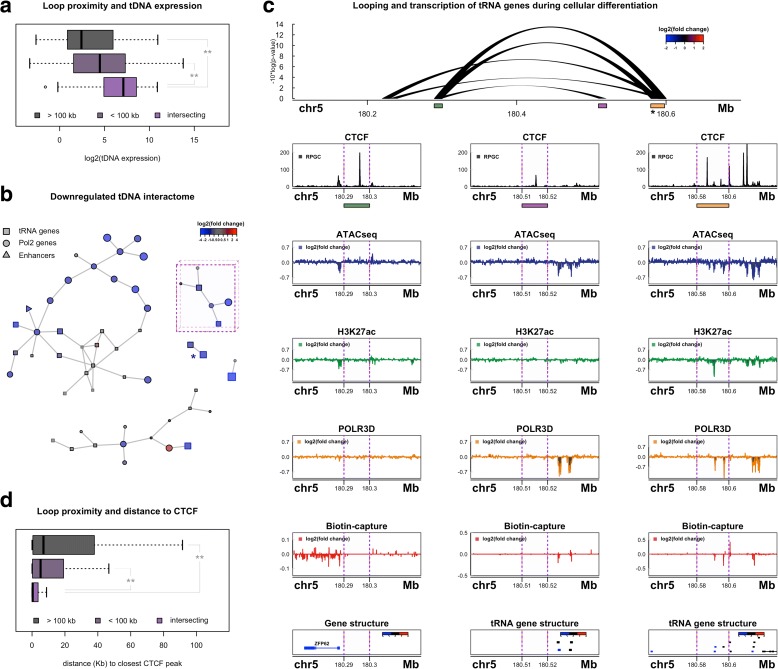
Coordinated long-range regulation of tRNA genes during cellular differentiation. a Distribution of integrated tDNA expression levels for tRNA genes > 100 Kb from a DNA loop end, within 100 Kb of a DNA loop end, and for tRNA genes that directly intersect DNA loop ends (left **p = 1.12^-6, right **p = 2.49^-11, Wilcoxon rank-sum test). b Network analysis of long-range interactions connecting tRNA genes downregulated in THP-1-derived macrophages. Each edge represents a DNA loop connecting two vertices (DNA loop anchors) that contain tRNA genes (square), RNAPII-transcribed genes (circle), or intergenic enhancers marked by H3K27 acetylation (triangle). Vertices with black frames represent loop anchors in which the identified feature (i.e. tRNA gene(s)) directly intersects the DNA loop end. Vertices without black frames represent loop anchors in which the identified feature is proximal to the DNA loop end (within 20 Kb). Both the size and color of each vertex is scaled by the mean log2(fold change) for resident feature(s). Purple outline marks the sub-community example further depicted in Fig. 4c. *Asterisk represents sub-community example further depicted in Additional file 1: Figure S5. c Visualization of chromatin and transcriptional dynamics at an example tDNA loop community located on chromosome 5. Colored rectangles define loop anchor regions further depicted below. *Asterisk represents loop anchor region depicted in Fig. 3a. Bottom left: signal track representation of CTCF binding sites (black, RPGC mean normalized reads per genomic content) and mean log2(fold change) for ATAC-seq (blue), H3K27ac (green), RNAPIII (orange), and nascent RNA (red) at the far-left loop anchor (green rectangle). Gene structure below includes RNAPII-transcribed gene ZFP62. Vertical dotted lines demarcate the actual loop anchor region. Bottom middle: analogous signal tracks depicting chromatin and transcriptional landscape at the middle loop anchor (purple rectangle) and proximal tDNA cluster. Bottom right: analogous signal tracks for the far-right loop anchor (orange rectangle) and intersecting tRNA genes. d Nearest distance to a CTCF binding site for tRNA genes that intersect DNA loop anchors, are within 100 Kb of a DNA loop anchor, or farther than 100 Kb from a DNA loop anchor (left **p = 2.31^-8, right **p = 6.61^-9, Wilcoxon rank-sum test)