Abstract
Background
The induction of cellulase production by insoluble carbon source cellulose was a common and efficient strategy, but has some drawbacks, such as difficult fermentation operation, substantial cellulase loss, long fermentation time, and high energy-consumption, resulting in high cost of cellulase production in industry. These drawbacks can be overcome if soluble carbon sources are utilized as the inducers for cellulase production. However, until now the induction efficiency of most soluble carbon sources, especially lactose and glucose, is still inferior to cellulose despite extensive efforts have been made by either optimizing the fermentation process or constructing the recombinant strains. Therefore, strain improvement by metabolic engineering for high induction efficiency of soluble carbon sources is of great interest.
Results
Trichoderma reesei mutant SEU-7 was constructed from T. reesei RUT-C30 with the overexpression of endogenous gene β-glucosidase (BGL1) by insertional mutagenesis via Agrobacterium tumefaciens-mediated transformation (AMT). Compared to RUT-C30, SEU-7 displays substantially enhanced activities of both cellulase and hemicellulase when grown on either lactose or cellulose. The induction efficiency with lactose was found to be higher than cellulose in strain SEU-7. To the best of our knowledge, we achieved the highest FPase activity in SEU-7 in both batch culture (13.0 IU/mL) and fed-batch culture (47.0 IU/mL) on lactose. Moreover, SEU-7 displayed unrivaled pNPGase activity on lactose in both batch culture (81.0 IU/mL) and fed-batch culture (144.0 IU/mL) as compared to the other reported T. reesei strains in the literature grown in batch or fed-batch experiments on cellulose or lactose. This superiority of SEU-7 over RUT-C30 improves markedly the saccharification ability of SEU-7 on pretreated corn stover. The overexpression of gene BGL1 was found either at the mRNA or at the protein level in the mutant strains with increased cellulase production in comparison with RUT-C30, but only SEU-7 displayed much higher expression of gene BGL1 on lactose than on cellulose. Two copies of gene BGL1 were inserted into the chromosome of T. reesei SEU-7 between KI911141.1:347357 and KI911141.1:347979, replacing the original 623-bp fragment that is not within any genes’ coding region. The qRT-PCR analysis revealed that the mRNA levels of both cellulase and hemicellulase were upregulated significantly in SEU-7, together with the MFS transporter CRT1 and the XYR1 nuclear importer KAP8.
Conclusions
Recombinant T. reesei SEU-7 displays hyper-production of both cellulase and hemicellulase on lactose with the highest FPase activity and pNPGase activity for T. reesei, enabling highly efficient saccharification of pretreated biomass. For the first time, the induction efficiency for cellulase production by lactose in T. reesei was reported to be higher than that by cellulose. This outperformance of T. reesei SEU-7, which is strain-specific, is attributed to both the overexpression of gene BGL and the collateral mutation. Moreover, the increased transcription levels of cellulase genes, the related transcription factors, and the MFS transporter CRT1 contribute to the outstanding cellulase production of SEU-7. Our research advances strain improvement to enhance the induction efficiency of soluble carbon sources to produce cost-effective cellulase and hemicellulase in industry.
Electronic supplementary material
The online version of this article (doi:10.1186/s13068-017-0915-9) contains supplementary material, which is available to authorized users.
Keywords: Trichoderma reesei, Soluble carbon source, Cellulase, Hemicellulase, β-Glucosidase, Hyper-production
Background
Although the insoluble cellulosic materials are considered as the most effective natural inducers for cellulase production by microorganisms in terms of both enzyme yield and productivity, their insolubility causes many problems and presents a major drawback that is partly responsible for the high cost of the cellulase production. First, the insolubility of cellulose leads to difficult and complex fermentation operations, including sterilization, cell biomass measurement, mixing and aeration of the fermentation broth [1], continuous feeding/sampling, and subsequent enzyme purification [2]. Second, cellulase gets absorbed to the solid cellulose surface, leading to enzyme loss. Third, cell growth and cellulase production are dependent on the cellulose hydrolysis, making the whole process time-consuming. Lastly, cellulase production induced by cellulose is very energy-intensive because of vigorous mixing and aeration of the viscous non-Newtonian fermentation broth [1]. These problems could be overcome if soluble inducers are used.
Previous studies showed that some soluble carbon sources alone can also induce the cellulase production, like sophorose [3, 4], cellobiose [5], galactose, l-sorbose [6], and lactose [7]. Sophorose is a strong inducer of cellulase, but it is very costly, and thus is not a practical carbon source for cellulase production [3]. The inducing effects of other soluble carbon sources are inferior to cellulose, partly due to central carbon catabolite repression (CCR).
Lactose, accumulated as the byproduct of cheese production, is inexpensive and economically feasible. Thus, it has become a major industrial inducer for (hemi)cellulolytic enzyme fermentation by T. reesei, though it is still less efficient in cellulase induction than cellulose owing to its slower induction rate and lower yield [5]. To improve the induction efficiency of lactose, several studies on strain improvement and fermentation process optimization have been carried out. For example, T. reesei mutant C-5 derived from QM9414 by mutagenesis with NTG can produce cellulase with 2.84 IU/mL FPase activity (Filter paper activity) on lactose [7], while overexpression of XYR1 under a copper responsive promoter in the ΔXYR1 strain derived from T. reesei QM9414 merely resulted in super low pNPCase (p-nitrophenyl-β-cellobioside) and pNPGase (p-n-nitrophenyl-β-glucosidase) activities on lactose [8]. The supplement of other carbon sources such as glucose [9] and cellulose + lactobionic acid [10], and different fermentation strategies like fed-batch culture [11] and continuous culture [9], have been utilized to help enhance the induction efficiency of lactose. These strategies are sometimes explored in combination to achieve a better induction effect. Despite the efforts shown above, the lactose induction efficiency is still low and not satisfying for cellulase industrial application.
In this research, T. reesei mutant SEU-7, derived from T. reesei RUT-C30 using insertional mutagenesis with AMT, displayed remarkably enhanced cellulase and hemicellulase production on both lactose and cellulose. For the first time, the cellulase production on lactose was reported to be superior to cellulose in T. reesei. The cellulase production of SEU-7 on lactose was further improved by optimization of lactose concentration, utilization of fed-batch culture, and supplement of low-concentration glucose, resulting in record FPase and pNPGase activities by T. reesei. Accordingly, improvement of the saccharification ability of SEU-7 cellulase on pretreated corn stover was observed. The role of the overexpression of gene BGL1 and the collateral mutation on the outperformance of SEU-7 were explored by profiling all other recombinants and determining the inserted copy numbers and the insertion sites of gene BGL1. The molecular mechanism for the outstanding performance of SEU-7 was further investigated by strain characterization and transcription analysis of the relevant genes.
Methods
Strains, plasmids, and culture media
Plasmid construction and propagation were carried out in Escherichia coli DH5α. A. tumefaciens AGL-1 was utilized as a T-DNA donor for the transformation of T. reesei RUT-C30 (CICC 13052) by A. tumefaciens-mediated transformation (AMT) [12]. E. coli DH5α and A. tumefaciens AGL-1 were grown in LB medium with 220 rpm at 37 and 28 °C, respectively. T. reesei was cultivated on potato dextrose agar (PDA) plate for conidia culture and in Trichoderma minimal media (TMM) [13] with 2% (w/t) cellulose or other carbon sources as indicated in the context for cellulase production at 28 °C with 200 rpm. The TMM medium was composed of the following (all concentrations in g/L unless otherwise noted): urea, 1.00; (NH4)2SO4, 4.00; KH2PO4, 6.59; FeSO4·7H2O, 0.005; MnSO4·H2O, 0.0016; ZnSO4·7H2O, 0.0014; CoCl2·6H2O, 0.002; MgSO4, 0.60; CaCl2, 0.60; Tween-80, 0.0186% (v/v); Tryptone, 0.75; Yeast extract, 0.25, Maleic acid, 11.6. The pH of TMM was adjusted to 6 by NaOH. Plasmid pDht/sk was a gift from Professor Zhihua Zhou from Key Laboratory of Synthetic Biology, Shanghai [14]. All chemicals used in this study were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Construction of recombinant T. reesei strains
By using HiScript 1st Strand cDNA Synthesis Kit (Vazyme, China), the first-strand cDNA was synthesized from total RNA of T. reesei RUT-C30 that was extracted with the RNA extraction Kit (Omega Bio-Tek, USA). BGL1 gene was amplified from T. reesei RUT-C30 cDNA with primers listed in Additional file 1: Table S1, and cloned into the backbone plasmid pDht/sk at XbaI using the ClonExpress II One Step Cloning Kit (Vazyme, China), generating plasmid pBGL-his (Additional file 1: Figure S1). Plasmid pBGL-his was transformed into T. reesei RUT-C30 by the AMT method [12]. After transferring the putative transformants on PDA medium containing hygromycin B successively for five generations, single spore colonies were isolated and five recombinant T. reesei strains were obtained: SEU-2, SEU-5, SEU-6, SEU-7, and SEU-8, in which the insertion of BGL1 gene was confirmed by PCR and sequencing.
Cellulase production
107/mL conidia produced by T. reesei SEU-7 or RUT-C30 grown on PDA plates for 7 days at 28 °C, which were counted by a Petroff-Hausser cell counter (American Optical, USA), were inoculated into 50-mL Erlenmeyer flasks containing 10 mL Sabouraud dextrose broth (SDB) and incubated for 48 h with 200 rpm at 28 °C. Pre-grown mycelia were inoculated with an inoculation ratio of 10% (v/v) into 250-mL flasks containing 50 mL TMM media (pH 6) with microcrystalline cellulose (Sinopharm Group Co. LTD), lactose, galactose, sucrose, cellobiose, glycerol, or glucose of different concentrations as shown in the context, and then incubated at 28 °C with 200 rpm. The fed-batch culture was carried out in the same way as the above batch culture except that 1.5 g autoclaved lactose powder, whose final concentration was 3%, was added at 120 h together with 5 mL 10 × TMM medium in one pulse. The volume of broth was 42 mL at the end of fermentation. When testing the influence of glucose supplementation on enzyme activities and protein concentration, 0.05–5% glucose was added into the TMM medium with 3% lactose or 2% cellulose as the inducer. Samples at different time points were taken for enzyme assay. Specifically, 1 mL culture was taken and centrifuged at 8000 rpm for 15 min at 4 °C to remove T. reesei cells and other solid materials, and the supernatant was stored at − 80 °C for further study. Each experiment was performed in three biological replicates.
Enzyme assay
The collected culture supernatant was diluted properly for enzyme assay. Enzyme activities of cellulase were measured by the standard protocols established in our previous study [15]. β-xylosidase activity was measured by using 4-nitrophenyl-β-d-xylopyranoside as a substrate dissolved in a 50 mM sodium citrate buffer, pH 5.0. The enzyme assay was performed at 50 °C as previously described [16]. Xylanase activity was determined with 1% beech xylan in 50 mM sodium acetate buffer as substrate [17]. 50 μL diluted culture supernatants were mixed with 50 μL substrate and reacted at 50 °C for 10 min. The reaction was stopped by adding 100 μL alkaline 3,5-dinitrosalicylic (DNS) and heating for 5 min at 100 °C. Then the absorbance at 540 nm was detected. Protein concentration was determined using BCA Protein Assay Kit (Sangon Biotech, China). All enzyme activities were presented as activities using international units (IU) per mL supernatant. One IU was defined as the amount of enzyme required to liberate 1 μmol of product per minute under the standard assay conditions. All experiments were performed in triplicate. The enzyme activities of the fed-batch experiment are calculated with the corrected broth volume to eliminate the concentration effect caused by evaporation.
Analysis method
Glucose was measured following the instructions of Glucose Detection Kit (Shanghai Rongsheng Biotech, China). For biomass measurement, 3 mL solid biomass was washed with tap water three times, and dried at 50 °C to constant weight [11]. DNA was determined as described in the literature [18]. SDS-PAGE analysis was performed on 10% Tris–HCl polyacrylamide gels using 10 μL cell culture of different T. reesei strains from day 7.
Saccharification of corn stover pretreated with EDA or alkali
Corn stover was harvested in suburb of Tianjin (China), air-dried, milled, and passed through a 2-mm sieve before pretreatment. The moisture of the milled corn stover was 4%. The milled corn stover was pretreated by ethylenediamine (PCS-EDA) in a vacuum drying oven as previously described [19, 20], which was termed as PCS-EDA. Corn stover containing 37.2% glucan and 19.2% xylan harvested in suburb of Jiangsu was air-dried, milled, and passed a 4-mm sieve with the moisture of 11% before pretreatment and treated by 2% dilute alkaline in a 1-L conical flask at 121 °C, 20 min [21], which was termed as PCS-alkali. Pretreated biomasses were then washed using water for several times until neutral pH. The washed biomasses were dried at 70 °C in an oven and stored at 4 °C before use.
The compositions of the pretreated corn stover (PCS) were determined following the Laboratory Analytical Procedure (LAP) of the National Renewable Energy Laboratory (NREL). We found that the PCS-EDA has 27.9% glucan, 9.9% xylan, and 16.6% acid-insoluble lignin in dry matter and the PCS-alkali 37.9% glucan, 19.2% xylan, and 16.7% acid-insoluble lignin. The other components in PCSs, such as soluble lignin, proteins, lipids, salts, were not analyzed here, leading to the PCS mass balances are well below 100%.
The supernatant collected at 168 h in batch culture and crude enzymes at 192 h in the fed-batch culture were used for saccharification. 10% (w/v) pretreated biomass in 1.5 mL buffer (50 mM sodium citrate buffer at pH 5.0 with 1 mM sodium azide to prevent microbial contamination) and different amount of crude enzyme were mixed and incubated at 50 °C with 400 rpm for 72 h. The reducing sugar level in the supernatant was determined by the DNS method. The experiments were performed in three biological repetitions.
Copy number assay
The copy numbers of the integrated gene BGL1 in the transformant strains SEU-2, 5, 6, 7, and 8 were determined by qPCR using extracted genomic DNA as the template and primers (Additional file 1: Table S2) on the basis of the reported method [22, 23]. Genomic DNAs of T. reesei strains were extracted individually with the E-Z 96 Fungal DNA Kits (Omega Bio-tek, Germany). Genes CEL7A, PGK1, and SAR1 were used to represent the single-copy genes, which is confirmed by nucleotide blasting against T. reesei genome sequence using the T. reesei genome database v2.0 (https://www.ncbi.nlm.nih.gov/genome/323?genome_assembly_id=49799). The qPCR was performed as follows: 95 °C for 30 s, 1 cycle of 95 °C for 15 s, 60 °C for 34 s, 40 cycles and 95 °C for 15 s, 60 °C for 60 s, 95 °C for 15 s 1 cycle. Three biological replicates were performed with three technical replicates for each biological replicate.
Whole genome resequencing
To identify the insert sites of gene BGL1 in the chromosome of T. reesei SEU-7, the genomes of SEU-7 and RUT-C30 were re-sequenced with Illumina HiSeq instrument according to manufacturer’s instructions (Illumina, San Diego, CA, USA). The NGS library was constructed following the manufacturer’s protocol (Illumina TruSeq DNA Nano Library Prep Kit). Then libraries with different indices were multiplexed and loaded on an Illumina HiSeq instrument according to manufacturer’s instructions (Illumina, San Diego, CA, USA). Sequencing was carried out using a 2 × 150 paired-end (PE) configuration; image analysis and base calling were conducted by the HiSeq Control Software (HCS) + OLB + GAPipeline-1.6 (Illumina) on the HiSeq instrument. The library construction and sequences were processed and analyzed by GENEWIZ (Su Zhou, China) using T. reesei RUT-C30 genome database (https://www.ncbi.nlm.nih.gov/genome/323?genome_assembly_id=49799). The corresponding fastq files of NGS for SEU-7 was deposited into NCBI SRA database with the NCBI Accession Number: SRR5931428.
qRT-PCR analysis
Reverse transcription (RT) was performed using the AceQ qPCR SYBR Green Master Mix (Takara, China) with total RNA that was prepared as described above from fresh mycelia of T. reesei SEU-7 and RUT-C30 cultivated under different conditions. The RNA concentration was determined at 260 nm using a NanoDrop ND-2000 (Thermo Fisher Scientific, USA). qRT-PCR was performed on the ABI StepOne instrument Plus (ABI, Germany) with software Version 2.3 (ABI, Germany). The primers for qRT-PCR are shown in Additional file 1: Table S3. At least three biological replicates were performed with three technical replicates for each biological replicate. The expression of PGK1 was chosen as the reference gene for data normalization [24].
Results
Recombinant T. reesei strain SEU-7 exhibits superior cellulase production ability for both cellulose and lactose to RUT-C30
The random insertion of a certain gene into a fungus’s chromosome for gene overexpression by the T-DNA-based mutagenesis usually interferes or disrupts other genes’ expression, leading to varied performance of different recombinant strains [5, 15, 24]. This phenomenon can be utilized as a powerful mutagenesis method in addition to the traditional ones using nitrosoguanidine (NTG) or UV irradiation. Here, endogenous gene β-glucosidase (BGL1, containing a linker and a C-terminal 6-histidine tag) was overexpressed in T. reesei under a modified CBH1 promoter [14] through the random insertion by AMT using the plasmid pBGL-his (Additional file 1: Figure S1A) to search for mutant strains with improved performance for cellulase production. Five transformants SEU-2, 5, 6, 7, and 8 were obtained with the cellulase production on cellulose assayed on day 5 (Additional file 1: Figure S1B). The pNPGase and CMCase (carboxymethyl cellulase) activities of all the tested recombinants were higher than the parental strain RUT-C30. However, the pNPCase activity of strains SEU-2 and SEU-5 was decreased, leading to the reduced FPase activity in these two strains. Although the strains SEU-6 and SEU-8 shared a similar pNPCase activity with RUT-C30, their FPase activity was lower than RUT-C30. Only the strain SEU-7 displayed increased pNPCase activity and FPase activity. Obviously, the strain SEU-7 with the highest cellulase activities outperforms other recombinant strains as well as the parent strain RUT-C30, and was therefore selected for further study.
Using cellulose as the sole carbon source, the FPase activity of the strain SEU-7 was 2.0-, 2.4-, and 2.2-fold that of RUT-C30 on days 3, 5, and 7, respectively (Fig. 1a). SEU-7 displayed 50 IU/mL pNPGase activity on day 7 (Fig. 1b), which was 34-fold that of RUT-C30. The pNPCase activity was increased by eightfold in SEU-7 on day 7 (Fig. 1c). The maximal CMCase activity of SEU-7 (21.3 IU/mL) was observed on day 5 and was 7.6-fold higher than that of RUT-C30 (2.8 IU/mL on day 7) (Fig. 1d). In agreement with the noticeable increment of cellulase activities, 42% more protein production was detected in the supernatant of SEU-7 cell culture (Fig. 1g). All together, SEU-7 outperforms the parent strain RUT-C30 by displaying markedly increment of both cellulase activities and secreted protein amount.
Fig. 1.
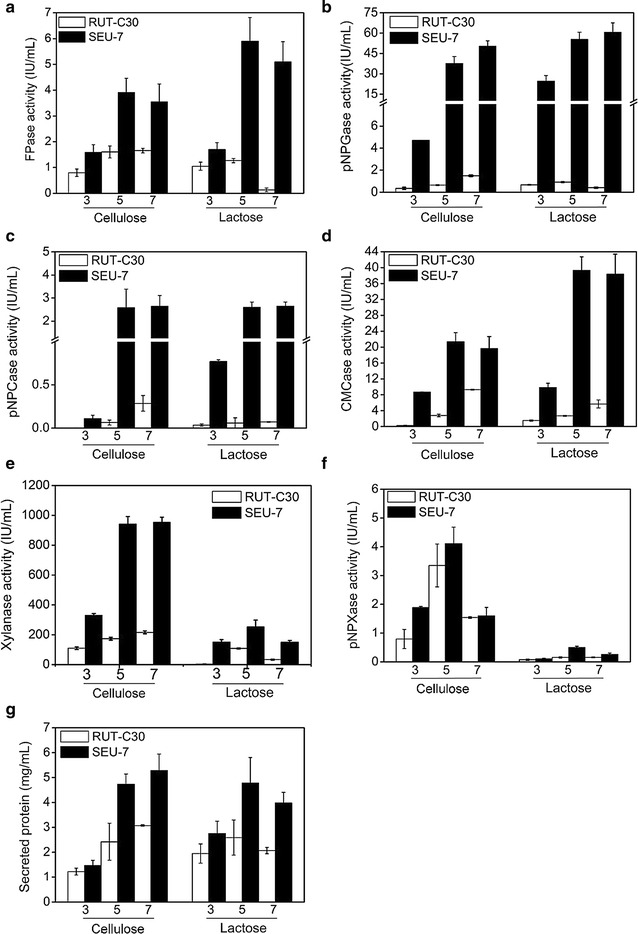
Cellulolytic enzyme activities in the culture supernatant of T. reesei SEU-7 and RUT-C30 grown on 2% cellulose or 2% lactose were assayed on day 3, day 5, and day 7, including the activities of FPase (the filter paper activity) (a), pNPGase (the BGL activity) (b), pNPCase (the CBH activity) (c), CMCase (the CMC activity) (d), xylanase (e) and pNPXase (the β-xylosidase activity) (f), and the protein concentration (g). The error bars indicate the standard deviations of three replicates
When grown on lactose, the maximal activities of FPase, pNPGase, pNPCase, and CMCase of SEU-7 were measured to be 5.9, 60.6, 2.6, and 39.0 IU/mL, respectively, which were 4.5-, 152.0-, 43.0- and 14.4-fold those of RUT-C30 on lactose, and were 3.7-, 40.6-, 43.0- and 14.0-fold those of RUT-C30 on cellulose (Fig. 1). As compared to those of SEU-7 grown on cellulose, the maximal activities of FPase, pNPGase, and CMCase of SEU-7 on lactose were improved by 51, 22, and 86%, respectively, while the pNPCase activity and the protein production were comparable. Obviously, the cellulase production of strain SEU-7 on 2% lactose is even better than that on 2% cellulose, which is in contrast to that result that the cellulase activities of RUT-C30 were compromised substantially on lactose as compared to cellulose (Fig. 1). The extracellular protein concentration of SEU-7 on lactose was also found to increase significantly 0.9-fold in comparison to RUT-C30 on lactose, demonstrating that the protein secretion ability of SEU-7 was improved. In summary, the induction efficiency of lactose was increased significantly in strain SEU-7, resulting in that the cellulase production of SEU-7 on lactose is higher than that of the industrial strain RUT-C30 on both cellulose and lactose.
SEU-7 exhibits better performance than RUT-C30 when grown on other soluble carbon sources including cellobiose, sucrose, galactose, glycerol, and glucose for cellulase production, but these carbon sources are not better cellulase inducers than cellulose or lactose for both strains (Additional file 1: Figure S2).
In addition, enhanced hemicellulase activity was also found in strain SEU-7 grown on either lactose or cellulose. The maximal xylanase activity of strain SEU-7 was measured to be 953 IU/mL on cellulose on day 7 and 253 IU/mL on lactose on day 5, which were 4.4- and 2.3-fold of those of RUT-C30, respectively (Fig. 1e). The final pNPXase (p-nitrophenyl-β-d-xylopyranoside) activity of SEU-7 was comparable to RUT-C30, although SEU-7 exhibited an increased pNPXase activity in the early phase day 3 (Fig. 1f).
The cellulase production of SEU-7 on lactose was further improved by lactose concentration optimization and fed-batch culture
It is reported that the cellulase production of T. reesei is dependent on the lactose content in the fermentation medium [7, 25, 26]. Therefore, the effect of lactose concentration on the cellulase production of strain SEU-7 was investigated (Fig. 2a). All the cellulase activities were increased gradually along with the increase of lactose concentration from 0.5 to 3%, and reached a plateau phase beyond 3% lactose, showing 3% lactose has the best induction effect for cellulase production in SEU-7. The FPase, pNPGase, pNPCase, CMCase, and pNPXase activities of SEU-7 induced by 3% lactose were 13.0, 81.0, 3.0, 62.0, and 0.5 IU/mL, respectively (Fig. 2a). When fed-batch culture with 3% lactose as the sole carbon source was explored, the maximum activities of FPase, pNPGase, and CMCase were 47.0, 140.0, and 164.0 IU/mL, respectively, at 192 h, while the maximal activities of pNPCase and pNPXase were 4.8 IU/mL at 168 h and 0.9 IU/mL at 144 h, respectively (Fig. 2c). The productivity and the yield of cellulase production were calculated using the FPase activity. When calculating the yield, it is assumed that lactose in the culture was metabolized completely as the lactose concentration was not measured. The FPase productivity was 0.10 and 0.24 IU/mL/h for the batch experiment and fed-batch experiment, repetitively, and the FPase yield individually 397 and 783 IU/g. The FPase productivity and yield was up 1.4- and 1.0-fold, respectively, by using fed-batch. The FPase productivity was higher than that of other previous studies with 0.2 [27] and 0.038 IU/mL/h [9] reported using lactose as inducer. Clearly, the cellulase production of SEU-7 on lactose was significantly enhanced by lactose concentration optimization and fed-batch culture.
Fig. 2.
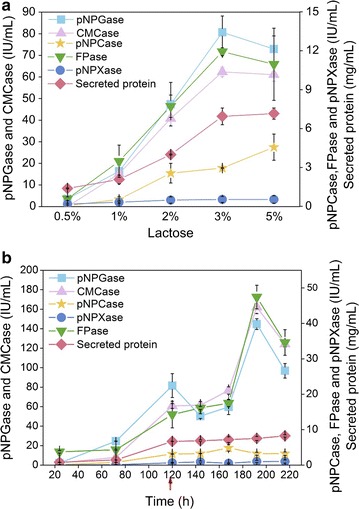
Cellulolytic enzyme activities of T. reesei SEU-7 and RUT-C30 grown on 0.5–5% lactose for 5 days (a) and cellulolytic enzyme activities of T. reesei SEU-7 in the fed-batch culture with 3% lactose (b). The error bars present the standard deviations of three biological replicates. The red arrow indicated the feeding time point
The cellulase production in strain SEU-7 was enhanced by the supplement of low-concentration glucose
As the sole carbon source in the medium, lactose serves as both an inducer for cellulase production and the carbon source for mycelium growth. The sharing of lactose for growth might limit the cellulase production. Given that strain SEU-7 displays a better resistance to CCR than RUT-C30 (Additional file 1: Figure S3), we envision that the supplement of glucose with low concentration for cell growth will not cause CCR, and will improve the cellulase production of SEU-7. Therefore, glucose in the range of 0–1% was added into the TMM medium containing 3% lactose and the cellulase activities were measured (Fig. 3a). The FPase activity kept rising from 13.0 to 20.0 IU/mL when the glucose concentration was increased from 0 to 0.1%, beyond which the FPase activity was decreased gradually. The pNPGase activity and the CMCase activity followed a similar trend as the FPase activity along with the increment of glucose concentration, peaking at 0.1% glucose. The maximal activities of pNPGase and CMCase were measured to be 90.0 and 78.0 IU/mL, respectively, which were improved by about 14.2 and 23.4%, respectively, when compared to those in the absence of glucose. The pNPCase activity stayed stable at 0–0.2% glucose, but dropped sharply when the glucose concentration used was > 0.2%. The protein concentration held steady until the glucose concentration was higher than 0.8% where it started to decline. In summary, the supplement of 0.1% glucose with 3% lactose has improved most cellulase production of SEU-7.
Fig. 3.
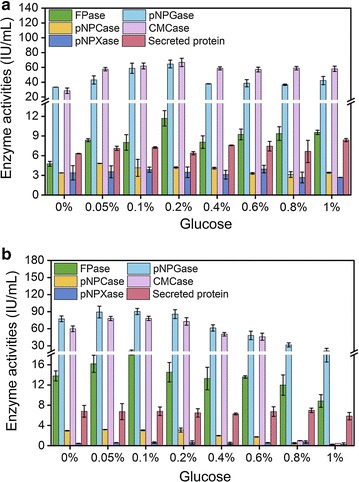
Effect of 0–1% glucose on cellulase production of strain SEU-7 grown on 3% lactose (a) or 2% cellulose (b). Cellulolytic enzyme activities were all measured on day 5. The error bars present the standard deviations of three biological tests
The benefit of the supplement of glucose was also observed with 2% cellulose as the carbon source. The maximal activities of FPase, pNPGase, and CMCase were found to be 12.0, 65.0, and 67.0 IU/mL, respectively, in the presence of 0.2% glucose (Fig. 3b). Furthermore, the pNPCase activity, pNPXase activity, and protein concentration were not changed as the glucose concentration varied. Apparently, the cellulase production of SEU-7 on lactose was more affected than on cellulose, when the mixed glucose concentration was increased from 0 to 1%, indicating that SEU-7 on lactose is more vulnerable to CCR than cellulose.
Characterization of T. reesei SEU-7 on lactose and cellulose
Although the strain SEU-7 displayed markedly improved cellulase activities on both lactose and cellulose than RUT-C30, its growth on lactose was noticeably slower than the strain RUT-C30 with 63% of the RUT-C30 biomass at 72 h but reached the same biomass as RUT-C30 at 96 h due to the steeply drop of the RUT-C30 biomass after 72 h (Fig. 4a). On cellulose, however, SEU-7 was grown better than RUT-C30 after 48 h with 1.5-fold more DNA content detected at 120 h (Fig. 4b). Due to the interference of insoluble cellulose to the measurement of dry T. reesei biomass, measurement of DNA content was used as an alternative method for biomass measurement.
Fig. 4.
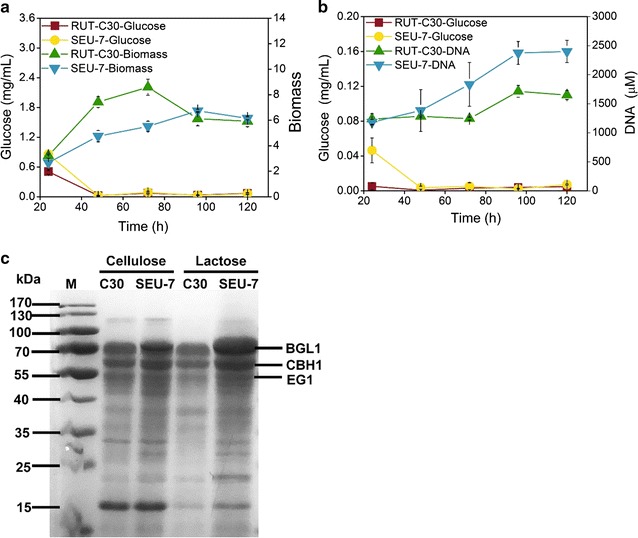
Growth and glucose concentration in the culture supernatant of T. reesei RUT-C30 and SEU-7 on 3% lactose (a) or 2% cellulose (b). SDS-PAGE analysis of secretome of RUT-C30 and SEU-7 grown on 3% lactose or 2% cellulose for 7 days (c)
69% more glucose was detected in the supernatant of SEU-7 than RUT-C30 on lactose at 24 h, after which the glucose concentration of both strains fell sharply to the identical concentration of 0.2 mg/mL at 48 h. It remained unaltered afterwards (Fig. 4a). When using cellulose as the carbon source, the strain SEU-7 had a very low glucose concentration (0.04 mg/mL) in its supernatant of 24 h, eightfold that of RUT-C30, which was followed by a sharply decline to 0.003 mg/mL at 48 h (Fig. 4b) and stayed unchanged in the rest of the fermentation process. In contrast, there was no obvious change of glucose concentration (~ 0.005 mg/mL) in the culture supernatant of RUT-C30 throughout the whole process of cellulase production (Fig. 4b). That more glucose was detected in the supernatant of SEU-7 culture on both lactose and cellulose might partly be attributed to significant enhancement of BGL activity found in SEU-7 on both carbon sources (Fig. 1b).
For both SEU-7 and RUT-C30, the glucose concentration is higher on lactose than on cellulose. This higher glucose concentration might cause more serious CCR, and partly contributes to the less efficient cellulase production of RUT-C30 on lactose. This did not happen to SEU-7 because of its relieved carbon catabolite depression: SEU-7 exhibited higher cellulase activities on glucose than RUT-C30 (Additional file 1: Figure S3).
The secretomes of T. reesei SEU-7 cultured in TMM with lactose or cellulose on day 7 were profiled by SDS-PAGE with equal volume of the culture supernatant (Fig. 4c). Obviously, SEU-7 secreted more cellulase than RUT-C30 into the culture medium, which is consistent with the observed increases of both cellulase activities (Fig. 1a) and protein concentration (Fig. 1g). It seems that SEU-7 possesses a better protein secretion ability than RUT-C30 when cultured on either lactose or cellulose.
The biomass saccharification ability of cellulase produced by strain SEU-7 was enhanced significantly
The hydrolysis efficiencies of crude enzyme solutions produced from the strain SEU-7 and RUT-C30 grown on lactose or cellulose, designated SEU-7-lac, SEU-7-cel, C30-lac, and C30-cel, respectively, were evaluated using corn stover pretreated with EDA (PCS-EDA) or alkali (PCS-alkali) as substrates (Fig. 5). On both substrates, both SEU-7-lac and SEU-7-cel produced much more reducing sugar than C30-lac or C30-cel at the same enzyme loading (Fig. 5). The maximal reducing sugar of SEU-7-lac (20.5 mg/mL with 80 μL enzyme loading) on PCS-EDA was five times that of C30-lac (4.1 mg/mL with 80 uL enzyme loading) (Fig. 5a). The maximal reducing sugar of SEU-7-lac (44.8 mg/mL with 40 μL enzyme loading) on PCS-alkali was fourfold that of C30-lac (11.2 mg/mL with 80 μL enzyme loading) (Fig. 5b). The hydrolysis efficiency of SEU-7-lac was a little lower than SEU-7-avi when using PCS-EDA or PCS-alkali as substrate. The hydrolysis efficiency of crude enzyme solution prepared from fed-batch culture of SEU-7 on lactose (SEU-7-lac-fed) was increased by 60 and 30% for PCS-EDA and PCS-alkali, respectively, as compared to SEU-7-lac. These results show that the mutant strain SEU-7 has much better biomass saccharification ability than RUT-C30 towards both EDA-pretreated biomass and alkali-pretreated biomass, enabling the significantly reduced usage amount of cellulase for hydrolysis of cellulose, which would help greatly reduce the cost of enzyme in industrial application.
Fig. 5.
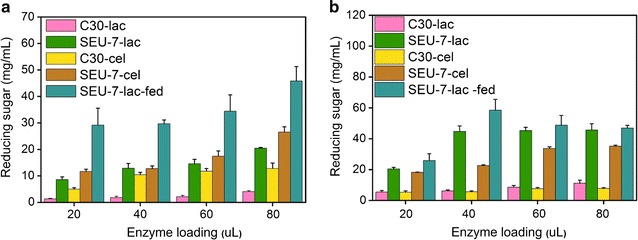
Saccharification of PCS with enzyme loading from 20 to 80 μL by T. reesei RUT-C30 and SEU-7 using the equal culture supernatants of RUT-C30 and SEU-7 on 2% cellulose or 3% lactose, respectively. Saccharification of corn stalk pretreated with EDA (a) or alkali (b) by crude enzyme solution produced from strain SEU-7 and RUT-C30 grown on lactose or cellulose, which was designated SEU-7-lac, SEU-7-cel, C30-lac, and C30-cel, respectively. The crude enzyme solution prepared from fed-batch culture of SEU-7 on 3% lactose was named as SEU-7-lac-fed. The error bars indicate the standard deviations of three biological replicates
The overexpression of gene BGL1 plays a positive role in the cellulase hyper-production of T. reesei SEU-7
To see how the overexpression of gene BGL1 impact the excellent performance of T. reesei SEU-7, the copy numbers and the mRNA level of gene BGL1, and the cellulase activities for all recombinants obtained in this study were determined. The copy numbers of gene BGL1 inserted were 2, 2, 1, 3, and 3 for strain SEU-7, SEU-2, SEU-5, SEU-6, and SEU-8, respectively, which were measured by qPCR (Fig. 6a). When grown on cellulose or lactose, strain SEU-7 and SEU-8 display a significant increase of BGL production at both the transcriptional level (Fig. 6b) and the protein level (Fig. 6c). Accordingly, SEU-7 and SEU-8 exhibited better cellulase production than RUT-C30 on both cellulose and lactose. A similar trend was observed for strain SEU-5 and SEU-6 on cellulose (Fig. 6c). Although their mRNA level of gene BGL1 was not enhanced on lactose, strain SEU-5 and SEU-6 display a remarkable increase in both BGL protein production and other cellulase production on lactose (Fig. 6c). The seemingly conflicting results between the mRNA expression level and the protein expression level are normal, since many other factors can influence protein expression such as the mRNA lifetime and the protein degradation rate. Conversely, the insertion of one copy of gene BGL1 into SEU-2 genome (Fig. 6a) did not result in the overexpression of gene BGL1 at either the mRNA level (Fig. 6b) or the protein level (Fig. 6c) when it was grown on cellulose or lactose. Meanwhile, SEU-2 exhibits similar cellulase production ability to RUT-C30 on both cellulose and lactose (Fig. 6c). It is worth mentioning that the mRNA level of gene BGL1 in strain SEU-7 on lactose was higher than on cellulose, which might contribute to the higher induction efficiency with lactose than with cellulose in strain SEU-7. On the contrary, it was reduced sharply in the other recombinants except strain SEU-2 on lactose as compared to cellulose. Obviously, the overexpression of gene BGL1 either at the mRNA or at the protein level plays a positive role in the increased cellulase production of T. reesei.
Fig. 6.
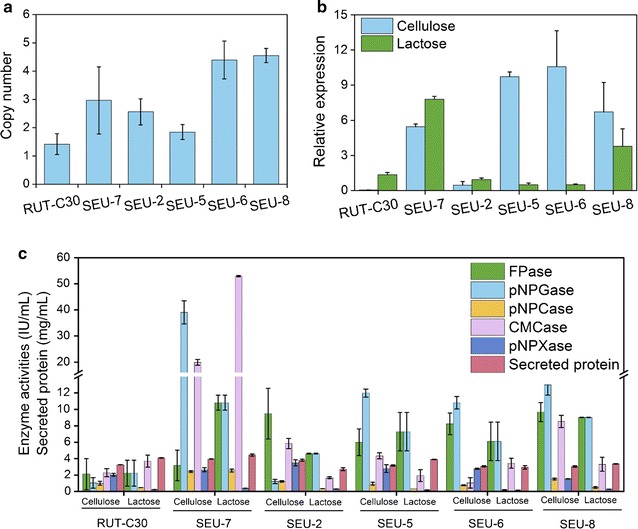
qPCR analysis of the copy numbers (a) and the transcript abundance (b) of BGL1 gene in T. reesei RUT-C30 and the five recombinant T. reesei strains: SEU-2, SEU-5, SEU-6, SEU-7, and SEU-8, and the cellulase activities of RUT-C30, SEU-7, SEU-2, SEU-5, SEU-6, and SEU-8 grown on cellulose and lactose for 5 days (c)
Identification of collateral mutation
The random insertion of gene BGL1 into the chromosome of T. reesei SEU-7 is possible to cause collateral mutations and interfere other genes’ expression, and in turn might contribute to the outstanding performance of SEU-7 together with the overexpression of gene BGL1. Hence, it would be impactful if insertional sites of the two inserted gene BGL1 are identified. To this end, the insertional sites were identified by the whole genome resequencing with NGS sequencing. The whole genome resequencing of T. reesei SEU-7 generated a total of 17849966 clean reads of 150-bp paired-end reads with mean depth coverage of 69.75 × and Q30 percentage of 88.69%. The obtained clean reads were mapped to the reference genome of RUT-C30, resulting in the high mean coverage of 99.41%. The NGS sequencing result (NCBI Accession Number: SRR5931428) show the two inserted copies of gene BGL1 was at 347357-347979 of T. reesei RUT-C30 genome KI911141.1 (https://www.ncbi.nlm.nih.gov/nuccore/572281258/), replacing the native fragment of 623 bp (Additional file 1: Figure S4). However, this replaced fragment was not within the coding region of any genes. The nearest downstream and upstream gene was M419DRAFT_127520 (KI911141.1:345112-346074) and M419DRAFT_6256 (KI911141.1:351617-351930), respectively, which is 1284 and 3639 bp away. Both genes are uncharacterized with unknown function. We tried to determine whether the expression of these two genes are affected by the replacement by measuring their mRNA levels with qRT-PCR, but failed because their transcriptional levels are too low to be detected under our current experimental conditions.
Transcription levels of (hemi)cellulase genes and regulators in T. reesei SEU-7
No matter the carbon source was either lactose or cellulose, the transcriptional levels of (hemi)cellulase genes β-glucosidase BGL1 (CEL3A), cellobiohydrolase CBHI (CEL7A), endoglucanase CMC (CEL7B), β-xylosidase BXL1, and xylanases (XYN1, XYN2, and XYN3) in strain SEU-7 were significantly higher than those of RUT-C30 (Fig. 7), matching well with the markedly enhanced (hemi)cellulase activities and increased (hemi)cellulase production amount found in the supernatant of SEU-7 culture.
Fig. 7.
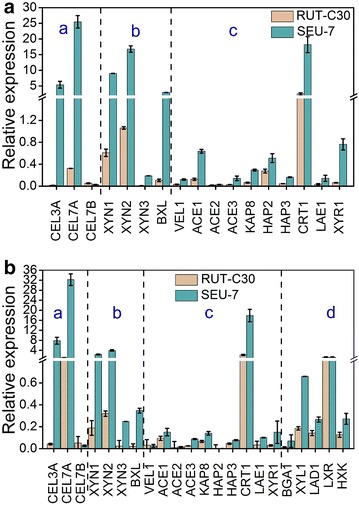
qRT-PCR analysis of the transcript abundance of genes relevant to cellulase and hemicellulase production in T. reesei SEU-7 and RUT-C30 grown on 2% cellulose (a) or 3% lactose (b) for 3 days. a: the main cellulase genes; b: the hemicellulase genes; c: regulatory genes involved in cellulase and hemicellulase production in T. reesei; d: genes participated in the lactose metabolism
Moreover, we analyzed the transcription of genes related to cellulase production, including cellulase transcription activators (XYR1, ACE2, ACE3, and HAP2/3/5), cellulase transcription repressors (ACE1 and CRE1), methyltransferase LAE1, nuclear importer KAP8, major facilitator superfamily (MFS) sugar transporter CRT1, β-glucosidase transcription factor BGLR, and global regulator VEL1(Fig. 7). Genes BGLR, HAP5, and CRE1 were not detected under our experiment conditions. On either lactose or cellulose, the mRNA expressions of both CRT1 and KAP8 were upregulated significantly in SEU-7, while those of LAE1 and VEL1 were unchanged. The transcription level of transcription activator ACE3 was enhanced in SEU-7 on lactose, but stayed constant in SEU-7 on cellulose. In contrast, the other transcription activators (XYR1, ACE2, and HAP2/3) showed constant mRNA levels in both SEU-7 and RUT-C30 on lactose, but displayed upregulated mRNA levels in SEU-7 on cellulose. Unexpectedly, the transcription level of transcription repressor ACE1 was markedly increased in SEU-7 on cellulose, which was not affected at all in SEU-7 on lactose.
The transcription levels of genes involved in lactose metabolism, including genes in both the Leroi pathway (GAL1, GAL7, and GAL10) and the alternative d-galactose pathways (BGA1, XYL1, XDH, HXK, LAD1, and LXR), were assayed in SEU-7 grown on lactose. Genes GAL1, BGA1, XYL1, LAD1, LXR, and HXK were detected. The mRNA levels of XYL1 was elevated remarkably in the strain SEU-7 in comparison to RUT-C30, while other detected genes’ transcription levels remained unchanged, indicating the capacity of these two pathways is abundant and is not a limitation that causes the inefficiency of lactose induction in T. reesei.
Discussion
Recently, several studies have been carried out in improving the induction efficiency of soluble carbon sources, mainly focusing on lactose and glucose that are cheap and abundant. Overexpression of XYR1 under a copper responsive promoter in the ΔXYR1 strain derived from T. reesei QM9414 merely resulted in super low pNPCase and pNPGase activities on glucose, lactose, and glycerol [8]. With constitutively overexpression of an artificial transcription activator including the XYR1 effectors, the XYR1 binding domains and the CRE1 binding domain in T. reesei RUT-C30, the recombinant strain zxy-2 can produce cellulase with 2.63 IU/mL FPase activity using glucose as a sole carbon source [28]. However, the cellulase production is incomplete with a very low pNPGase activity, leading to the requirement for pNPGase supplement during the saccharification [28]. A sophorose-containing soluble sugar mixture prepared by the transglycosylation reaction catalyzed by β-glucosidase was investigated to obtain an effective synthetic sugar mixture, which can achieve a cellulase titer of 90.3 FPU (filter paper unit)/mL in the fed-batch fermentation of T. reesei RUT-C30 on glucose [11]. Nevertheless, this strategy makes the cellulase production process more complex and inconvenient, and introduces extra cost by using the β-glucosidase to carry out the transglycosylation. In addition, the heavy reliance of cellulase production on substrate composition is not flexible and not economic feasible for applications in industry at a large scale. Noticeably, up to now, there are several issues in cellulase production by insoluble carbon sources, including the low productivity, the incomplete compositions of the cellulase production, the extra addition of β-glucosidase for transglycosylation reaction. These issues prevent the economic cellulase production.
Here, T. reesei SEU-7 constructed from T. reesei RUT-C30 using the T-DNA-based mutagenesis outperforms RUT-C30 in terms of cellulase production on either cellulose or lactose. Most importantly, T. reesei SEU-7 is the first T. reesei mutant ever reported which cellulase production ability on lactose is better than that on cellulose. The cellulase production of SEU-7 on lactose was further improved by the optimization of lactose concentration, utilization of fed-batch culture, and supplement of low-concentration glucose, resulting in record FPase and pNPGase activities. For SEU-7, the FPase activity of 13.0 IU/mL by batch culture and 47.0 IU/mL by fed-batch culture were 4.6- and 16.5-fold those ever reported (the highest reported FPase activity was 2.84 IU/mL) in T. reesei when using lactose as the sole carbon source [8]. To the best of our knowledge, these FPase activities of T. reesei SEU-7 when using lactose as the sole carbon source are the highest. Moreover, the pNPGase activity of 81 IU/mL by batch culture and 140.0 IU/mL by fed-batch culture were all higher than the highest BGL activity of 69.7 IU/mL ever reported in T. reesei grown on cellulose [22]. This solves the dwindling issue of low β-glucosidase activity in T. reesei cellulase that leads to inefficiency in biomass degradation and limits its industrial application [29, 30]. Even better, the pNPCase activity, CMCase activity, and FPase activity of SEU-7 on 3% lactose were all higher than strain T4 on cellulose [22].
Moreover, it is worth mentioning that a complete cellulase production was obtained for SEU-7. Complete enzyme production of SEU-7 enables highly effective hydrolysis of not only EDA-pretreated biomass, but also alkali-pretreated biomass that contains more xylan [11], demonstrating that it allows a broad choice of substrates for saccharification. Meanwhile, improvement of the saccharification ability of SEU-7 compared to RUT-C30 on pretreated corn stover was observed (Fig. 5). These merits make SEU-7 a preferable producer that will work efficiently in the industrial application when cellulose or lactose is utilized as the carbon source for cellulase production.
The benefit of the supplement of low-concentration glucose was observed when using 3% lactose or 2% cellulose as the carbon source, which is probably due to both the relieved CCR and the significantly increased production of β-glucosidase in strain SEU-7. Since strain SEU-7 displays a better resistance to CCR than RUT-C30 (Additional file 1: Figure S3), the supplement of glucose with low concentration did not cause CCR. Instead, the added glucose can serve as the extra carbon source for mycelium growth, saving more lactose or cellulose as the inducer for cellulase production. Moreover, glucose could be transglycosylated by the increased β-glucosidase into sophorose that is a strong inducer for cellulase production [11].
In our opinion, two factors might be responsible for the protein concentration being almost constant, whereas FPase and CMCase activities are multiplied by 2 or 3 after 120 h in the fed-batch experiment (Fig. 2b). Besides cellulase, the secreted protein complex contained non-cellulase proteins like pectinase, mannanase, and the auxiliary proteins which have been continuously found to help improve the hydrolysis of cellulose, such as swollenin (encoded by swo1) [31], the Glycoside Hydrolases Family 61 (GH61s) [32], and the cellulase-induced proteins CIP-1 and CIP-2 [33]. No increase in the content of the secreted protein after 120 h does not mean that the amount of cellulase is not increased either. On the other hand, only the three major cellulolytic components, CBH, CMC, and BGL, are not enough for high efficiency of the cellulose hydrolysis, which was demonstrated by the low cellulose hydrolysis yield from the reconstituted mixture consisting of CBH, CMC, and BGL [34]. Low abundant essential enzymes in the supernatant of T. reesei culture are also important for the high efficiency of T. reesei cellulase. Thus, even if the amount of cellulase is not increased after 120 h, the FPase activity might be improved due to the elevated expression of these low abundant essential enzymes, which is too low to be detected by our protein concentration method.
Genetic engineering has been extensively exploited to increase the BGL activity in T. reesei, sometimes as well as the other cellulase compositions, by the overexpression of the native β-glucosidase BGL1 [15, 35] as done in this study or the exotic ones from other fungi including Neosartorya fischeri [22], Penicillium decumbens [36], Aspergillus aculeatus [29, 30], and Periconia sp. [37]. Among all these mutant strains, only strain TRB1 has been grown on lactose for cellulase production that is declined in comparison with cellulose [15]. The cellulase production of other recombinants with the overexpression of gene BGL1 in this study like SEU-5, SEU-7, and SEU-8 is more or less compromised on lactose than on cellulase. Moreover, along with the overexpression of BGL1, one or several other cellulase compositions were increased [15, 22, 29, 30, 35, 36] as found in the recombinants SEU-5, SEU-7, and SEU-8 here, but it is rare to get all other cellulase compositions increase as we observed in strain SEU-7. Therefore, the outperformance of SEU-7 on both cellulose and lactose is strain-specific, which might be owing to the fine-tuning of the BGL1 expression to achieve an optimal level, and/or the collateral mutations caused by random insertion.
Two factors might be involved in the role of the BGL1 overexpression playing in the outperformance of strain SEU-7 for cellulase production. The synthetic overexpression of BGL1 both at the mRNA level and at the protein level in strain SEU-7 might play a role in the altered regulatory effect found in strain SEU-7 by transcriptional analysis, which is beneficial for the notable increment of cellulase activities and the overall secreted protein. Besides this, the noticeably improved overexpression of gene BGL1 can increase the sophorose concentration to have more strong inducer for strain SEU-7 to produce cellulase production, for protein BGL1 can catalyze the conversion of glucose and cellobiose to sophorose that is strong inducers for cellulase production [11, 38].
The random insertion of a gene into chromosome for overexpression is possible to interfere other genes’ expression, and in turn affects the performance of T. reesei in addition to the overexpression of the gene. As a result, different recombinant strains with the overexpression of the same gene might display varied performance, which was observed in our study. However, this effect in most cases has not been taken into account for the performance of the mutant strains in previous studies [30, 37]. The effect of the insertion position should definitely be considered when dealing with gene modification by random integration. The insertion of two copies of gene BGL1 into the chromosome of T. reesei SEU-7 between KI911141.1:347357 and KI911141.1:347979 has caused a collateral mutation, the deletion of the original 623-bp fragment. Nevertheless, how this collateral mutation interferes other genes in SEU-7 is unknown, for the deleted gene sequence was not part of any open-reading frame (ORFs) of genes, whereas the mRNA level of its two adjacent genes M419DRAFT_127520 and M419DRAFT_6256 is under the limitation of detection by RT-qPCR. The missing of the original 623-bp fragment might take a part in the outperformance of SEU-7 at the chromatin-level regulation through chromatin remodeling as reported for the transcription factors involved in cellulase production. For example, both the transcriptional activator XYR1 and the transcriptional repressor CRE1 are involved in the chromatin remodeling. Chromatin rearrangement was observed in the XYR1 promoter during induction on sophorose, and the XYR1 promoter is overall more accessible in a truncated CRE1 background [39]. Also, CRE1 plays a key role in chromatin packaging under repression conditions [40–42]. CRE1 and the HAP2/3/5 complex take part in nucleosome assembly in the CBH2 promoter [42]. Furthermore, the insertion position probably affects the expression of the two inserted copies of BGL1 that is important as we have discussed above. Nevertheless, further investigations are required to explore these possibilities.
Transcription analysis of genes encoding major cellulases and the related genes with cellulase production were performed to identify key factors contributing to the excellent performance of SEU-7 over RUT-C30. Along with their enhanced protein expression and enzyme activities, significant upregulation was observed with the mRNA expressions of (hemi)cellulase genes. The MFS transporter CRT1 (tre3405) in T. reesei, which is considered as lactose permease [43, 44], is essential for cellulase production on cellulose, lactose, or sophorose, and is highly expressed during the growth of T. reesei on cellulose [45], lactose [45], and wheat straw [46]. These previous results well explain the remarkably elevated mRNA level of CRT1 we observed in SEU-7. It is reported that CRT1 was downregulated (78-fold) in the mutant strain ∆XYR1 [47] when cellulose was used as the carbon source, suggesting a positive correlation between XYR1 and CRT1 occurs in the regulation of cellulase production. Consistently, the transcriptional level of XYR1 was unregulated in SEU-7 on cellulose, which was accompanied with the surged mRNA level of KAP8 that mediates the nuclear import of protein XYR1 [43]. Meanwhile, activator ACE3 displayed an enhanced mRNA level in SEU-7 on lactose, while activators ACE2 and HAP2/3 showed upregulated mRNA levels in SEU-7 on cellulose. On the contrary, the mRNA levels of genes participating in lactose metabolism were not changed noticeably in SEU-7. It seems that the increased transcription levels of cellulase genes, the related transcription factors, and the MFS transporter CRT1 contribute to the outstanding cellulase production of SEU-7.
Unlike model microorganisms E. coli and yeast, T. reesei does not possess well-developed genetic tools. Therefore, its engineering relies heavily on NTG mutagenesis, UV mutagenesis, and knockout or overexpression of a certain well-known gene to get mutant strains with desired performance, which definitely limits the ability to get T. reesei mutant strains with desired performance. Furthermore, the mechanism of cellulase production is unknown, preventing strain engineering with rational design. Thus, the random insertion method of AMT presents a useful strategy, as demonstrated here and in the previous studies [15, 48, 49], but has not been fully utilized for strain engineering in T. reesei and other fungi.
Conclusions
It is the first time to obtain a T. reesei mutant SEU-7 that enables the soluble carbon source lactose as an efficient inducer as cellulose, presenting a substantial progress in the utilization of soluble carbon sources for cellulase production in T. reesei. In addition to the high yield, SEU-7 produced a complete whole set of cellulase and hemicellulase that are all improved notably on both cellulose and lactose, allowing a broad choice of pretreated biomass for saccharification. SEU-7 displays record FPase activity and BGL activity when grown on lactose. This outperformance of T. reesei SEU-7 is strain-specific as demonstrated by its different overexpression pattern of gene BGL1 from other recombinants, and is attributed to both the elevated expression level of gene BGL1 and the collateral mutation. Two copies of the inserted gene BGL1 were detected at the chromosome of T. reesei SEU-7 between KI911141.1:347357 and KI911141.1:347979, deleting the original 623-bp fragment that is not within any genes’ coding region. Unfortunately, it is still unclear how the identified collateral mutation gets involved in the excellent performance of SEU-7. Strain SEU-7 was altered significantly at gene transcription level as well as its phenotype like growth, extracellular glucose concentration, and protein secretion ability. T. reesei SEU-7 is definitely a better alternative to RUT-C30 in industry for cost-effective cellulase production using either lactose or cellulose.
Authors’ contributions
CL and FL conceived and designed the study. CL carried out the majority of the experiments. LZ conducted part of enzyme activity measurement experiments. ZZ provided the plasmid pDht/sk and helped us to design and construct the plasmid pBGL-his used in this study. LQ and BL conducted the EDA pretreatment of corn stover. MJ prepared the alkali-pretreated corn stover and analyzed its compositions. CL, FL, and ZC analyzed the data and drafted the manuscript. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Availability of supporting data
Additional file 1.
Consent for publication
Not applicable.
Ethical approval and consent to participate
Not applicable.
Funding
This work was supported by the Fundamental Research Funds for the Central Universities, a project funded by the Priority Academic Program Development (PAPD) of Jiangsu Higher Education Institutions, and the scientific research innovation projects for ordinary university academic degree graduate student of Jiangsu province (KYLX16-0290). ZC thanks the University of Michigan for supporting his sabbatical leave.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- pNPGase
the β-glucosidase activity
- pNPCase
the CBH activity
- CMCase
the CMC activity
- FPase
the filter paper activity
- pNPXase
the β-xylosidase activity
- AMT
agrobacterium tumefaciens-mediated transformation
- PDA
potato dextrose agar
- SDB
sabouraud dextrose broth
- TMM
Trichoderma minimal medium
- IU
international units
- pNPG
p-nitrophenyl-β-d-glucopyranoside
- EDA
ethylenediamine
- PCS
the pretreated corn stover
- DNS
3,5-dinitrosalicylic
- CCR
carbon catabolite repression
- ORF
open-reading frame
- DNS
3,5-dinitrosalicylic
- RT
reverse transcription
- MFS
major facilitator superfamily sugar transporter
Additional file
Additional file 1: Figure S1. Schematic illustration of the plasmid pBGL. Figure S2. Cellulase activities of SEU-7 and RUT-C30 grown on different carbon sources. Figure S3. Effect of glucose on the enzyme activities of T. reesei RUT-C30 and SEU-7. Figure S4. The DNA sequence of the missing fragment at the loci of KI911141.1:351617-351930. Table S1. PCR primers for plasmid construction. Table S2. Primers for determination of copy numbers of BGL1 in SEU-7 using qPCR. Table S3. Primers for real-time quantitative PCR.
Footnotes
Electronic supplementary material
The online version of this article (doi:10.1186/s13068-017-0915-9) contains supplementary material, which is available to authorized users.
Contributor Information
Chengcheng Li, Email: lichengcheng1162@126.com.
Fengming Lin, Phone: +86-83791810, Email: linfengming@seu.edu.cn.
Le Zhou, Email: 2817055928@qq.com.
Lei Qin, Email: qinlei@tju.edu.cn.
Bingzhi Li, Email: bzli@tju.edu.cn.
Zhihua Zhou, Email: zhouzhihua@sippe.ac.cn.
Mingjie Jin, Email: jinmingjie@njust.edu.cn.
Zhan Chen, Email: zhanc2010@gmail.com.
References
- 1.Gabelle JC, et al. Impact of rheology on the mass transfer coefficient during the growth phase of Trichoderma reesei in stirred bioreactors. Chem Eng Sci. 2012;75:408–417. doi: 10.1016/j.ces.2012.03.053. [DOI] [Google Scholar]
- 2.Reese ET, Mandeis M. Production of microbial enzymes for cellulose hydrolysis. Ciencia biologica Mol Cell Biol. 1980;5:11. [Google Scholar]
- 3.Mach RL, et al. The bgl1 gene of Trichoderma reesei QM 9414 encodes an extracellular, cellulose-inducible beta-glucosidase involved in cellulase induction by sophorose. Mol Microbiol. 1995;16:687–697. doi: 10.1111/j.1365-2958.1995.tb02430.x. [DOI] [PubMed] [Google Scholar]
- 4.Mandels M, Parrish FW, Reese ET. Sophorose as an inducer of cellulase in Trichoderma viride. J Bacteriol. 1962;83:400–408. doi: 10.1128/jb.83.2.400-408.1962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Warzywoda M, Ferre V, Pourquie J. Development of a culture medium for large-scale production of cellulolytic enzymes by Trichoderma reesei. Biotechnol Bioeng. 1983;25:3005–3011. doi: 10.1002/bit.260251216. [DOI] [PubMed] [Google Scholar]
- 6.Kawamori M, Morikawa Y, Takasawa S. Induction and production of cellulases by l-sorbose in Trichoderma reesei. Appl Microbiol Biotechnol. 1986;24:449–453. [Google Scholar]
- 7.Chaudhuri B, Sahai V. Production of cellulase using a mutant strain of Trichoderma reesei growing on lactose in batch culture. Appl Microbiol Biotechnol. 1993;39:194–196. doi: 10.1007/BF00228605. [DOI] [Google Scholar]
- 8.Lv X, Zheng F, Li C, et al. Characterization of a copper responsive promoter and its mediated overexpression of the xylanase regulator 1 results in an induction-independent production of cellulases in Trichoderma reesei. Biotechnol Biofuels. 2015;8:67. doi: 10.1186/s13068-015-0249-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Domingues FC, Queiroz JA, Cabral JMS, et al. Production of cellulases in batch culture using a mutant strain of Trichoderma reesei growing on soluble carbon source. Biotechnol Lett. 2001;23:771–775. doi: 10.1023/A:1010329731010. [DOI] [Google Scholar]
- 10.Ahamed A, Vermette P. Culture-based strategies to enhance cellulase enzyme production from Trichoderma reesei RUT-C30 in bioreactor culture conditions. Biochem Eng J. 2008;40:399–407. doi: 10.1016/j.bej.2007.11.030. [DOI] [Google Scholar]
- 11.Li Y, Liu C, Bai F, et al. Overproduction of cellulase by Trichoderma reesei RUT C30 through batch-feeding of synthesized low-cost sugar mixture. Bioresour Technol. 2016;216:503–510. doi: 10.1016/j.biortech.2016.05.108. [DOI] [PubMed] [Google Scholar]
- 12.Zhong YH, Wang XL, Wang TH, et al. Agrobacterium-mediated transformation (AMT) of Trichoderma reesei as an efficient tool for random insertional mutagenesis. Appl Microbiol Biotechnol. 2007;73:1348–1354. doi: 10.1007/s00253-006-0603-3. [DOI] [PubMed] [Google Scholar]
- 13.Minty JJ, Singer ME, Scholz SA, et al. Design and characterization of synthetic fungal-bacterial consortia for direct production of isobutanol from cellulosic biomass. Proc Natl Acad Sci USA. 2013;110:14592–14597. doi: 10.1073/pnas.1218447110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zou G, Shi S, Jiang Y, et al. Construction of a cellulase hyper-expression system in Trichoderma reesei by promoter and enzyme engineering. Microb Cell Fact. 2012;11:21. doi: 10.1186/1475-2859-11-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li C, et al. A beta-glucosidase hyper-production Trichoderma reesei mutant reveals a potential role of cel3D in cellulase production. Microb Cell Fact. 2016;15:151. doi: 10.1186/s12934-016-0550-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kristufek D, Zeilinger S, Kubicek CP. Regulation of β-xylosidase formation by xylose in Trichoderma reesei. Appl Microbiol Biotechnol. 1995;42:713–717. doi: 10.1007/BF00171950. [DOI] [Google Scholar]
- 17.Törrönen A, Mach RL, Messner R, et al. The two major xylanases from Trichoderma reesei: characterization of both enzymes and genes. Nat Biotechnol. 1992;10:1461–1465. doi: 10.1038/nbt1192-1461. [DOI] [PubMed] [Google Scholar]
- 18.Wang M, Zhao Q, Yang J, et al. A mitogen-activated protein kinase Tmk3 participates in high osmolarity resistance, cell wall integrity maintenance and cellulase production regulation in Trichoderma reesei. PLoS ONE. 2013;8:e72189. doi: 10.1371/journal.pone.0072189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Qin L, Li WC, Zhu JQ, et al. Ethylenediamine pretreatment changes cellulose allomorph and lignin structure of lignocellulose at ambient pressure. Biotechnol Biofuels. 2015;8:174. doi: 10.1186/s13068-015-0359-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Qin L, Liu L, Li WC, et al. Evaluation of soluble fraction and enzymatic residual fraction of dilute dry acid, ethylenediamine, and steam explosion pretreated corn stover on the enzymatic hydrolysis of cellulose. Bioresour Technol. 2016;209:172–179. doi: 10.1016/j.biortech.2016.02.123. [DOI] [PubMed] [Google Scholar]
- 21.McIntosh S, Vancov T. Optimisation of dilute alkaline pretreatment for enzymatic saccharification of wheat straw. Biomass Bioenergy. 2011;35:3094–3103. doi: 10.1016/j.biombioe.2011.04.018. [DOI] [Google Scholar]
- 22.Xue X, et al. Revisiting overexpression of a heterologous β-glucosidase in Trichoderma reesei: fusion expression of the Neosartorya fischeri Bgl3A to cbh1 enhances the overall as well as individual cellulase activities. Microb Cell Fact. 2016;15:122. doi: 10.1186/s12934-016-0520-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Solomon PS, Ipcho SVS, Hane JK, et al. A quantitative PCR approach to determine gene copy number. Fungal Genet Rep. 2008;55:5–8. doi: 10.4148/1941-4765.1082. [DOI] [Google Scholar]
- 24.Rahman Z, Shida Y, Furukawa T, et al. Application of Trichoderma reesei cellulase and xylanase promoters through homologous recombination for enhanced production of extracellular β-glucosidase I. Biosci Biotechnol Biochem. 2009;73:1083–1089. doi: 10.1271/bbb.80852. [DOI] [PubMed] [Google Scholar]
- 25.Sehnem NT, de Bittencourt LR, Camassola M, et al. Cellulase production by Penicillium echinulatum on lactose. Appl Microbiol Biotechnol. 2006;72(1):163–167. doi: 10.1007/s00253-005-0251-z. [DOI] [PubMed] [Google Scholar]
- 26.Morikawa Y, Ohashi T, Mantani O, et al. Cellulase induction by lactose in Trichoderma reesei PC-3-7. Appl Microbiol Biotechnol. 1995;44:106–111. doi: 10.1007/BF00164488. [DOI] [Google Scholar]
- 27.Ahamed A, Vermette P. Effect of mechanical agitation on the production of cellulases by Trichoderma reesei RUT-C30 in a draft-tube airlift bioreactor. Biochem Eng J. 2010;49:379–387. doi: 10.1016/j.bej.2010.01.014. [DOI] [Google Scholar]
- 28.Zhang X, Li Y, Zhao X, et al. Constitutive cellulase production from glucose using the recombinant Trichoderma reesei strain overexpressing an artificial transcription activator. Bioresour Technol. 2017;223:317–322. doi: 10.1016/j.biortech.2016.10.083. [DOI] [PubMed] [Google Scholar]
- 29.Nakazawa H, et al. Construction of a recombinant Trichoderma reesei strain expressing Aspergillus aculeatus beta-glucosidase 1 for efficient biomass conversion. Biotechnol Bioeng. 2012;109:92–99. doi: 10.1002/bit.23296. [DOI] [PubMed] [Google Scholar]
- 30.Treebupachatsakul T, et al. Utilization of recombinant Trichoderma reesei expressing Aspergillus aculeatus beta-glucosidase I (JN11) for a more economical production of ethanol from lignocellulosic biomass. J Biosci Bioeng. 2015;120:657–665. doi: 10.1016/j.jbiosc.2015.04.015. [DOI] [PubMed] [Google Scholar]
- 31.Saloheimo M, et al. Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur J Biochem. 2002;269:4202–4211. doi: 10.1046/j.1432-1033.2002.03095.x. [DOI] [PubMed] [Google Scholar]
- 32.Merino ST, Cherry J. Progress and challenges in enzyme development for biomass utilization. In: Olsson L, editor. Biofuels. Berlin: Springer; 2007. pp. 95–120. [DOI] [PubMed] [Google Scholar]
- 33.Kuhad RC, et al. Revisiting cellulase production and redefining current strategies based on major challenges. Renew Sustain Energy Rev. 2016;55:249–272. doi: 10.1016/j.rser.2015.10.132. [DOI] [Google Scholar]
- 34.Adav SS, Chao LT, Sze SK. Quantitative secretomic analysis of Trichoderma reesei strains reveals enzymatic composition for lignocellulosic biomass degradation. Mol Cell Proteom. 2012;11(M111):012419. doi: 10.1074/mcp.M111.012419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhang J, Zhong Y, Zhao X, et al. Development of the cellulolytic fungus Trichoderma reesei strain with enhanced β-glucosidase and filter paper activity using strong artifical cellobiohydrolase 1 promoter. Bioresour Technol. 2010;101:9815–9818. doi: 10.1016/j.biortech.2010.07.078. [DOI] [PubMed] [Google Scholar]
- 36.Ma L, et al. Improvement of cellulase activity in Trichoderma reesei by heterologous expression of a beta-glucosidase gene from Penicillium decumbens. Enzyme Microb Technol. 2011;49:366–371. doi: 10.1016/j.enzmictec.2011.06.013. [DOI] [PubMed] [Google Scholar]
- 37.Dashtban M, Qin W. Overexpression of an exotic thermotolerant β-glucosidase in Trichoderma reesei and its significant increase in cellulolytic activity and saccharification of barley straw. Microb Cell Fact. 2012;11:63. doi: 10.1186/1475-2859-11-63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Guo B, Sato N, Biely P, et al. Comparison of catalytic properties of multiple β-glucosidases of Trichoderma reesei. Appl Microbiol Biotechnol. 2016;100:4959–4968. doi: 10.1007/s00253-016-7342-x. [DOI] [PubMed] [Google Scholar]
- 39.Mello-de-Sousa TM, et al. The relation between promoter chromatin status, Xyr1 and cellulase expression in Trichoderma reesei. Curr Genom. 2016;17:145–152. doi: 10.2174/1389202917666151116211812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mello-de-Sousa TM, et al. A truncated form of the carbon catabolite repressor 1 increases cellulase production in Trichoderma reesei. Biotechnol Biofuels. 2014;7:129. doi: 10.1186/s13068-014-0129-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ries L, et al. The role of CRE1 in nucleosome positioning within the cbh1 promoter and coding regions of Trichoderma reesei. Appl Microbiol Biotechnol. 2014;98:749–762. doi: 10.1007/s00253-013-5354-3. [DOI] [PubMed] [Google Scholar]
- 42.Zeilinger S, et al. Nucleosome transactions on the Hypocrea jecorina (Trichoderma reesei) cellulase promoter cbh2 associated with cellulase induction. Mol Genet Genom. 2003;270:46–55. doi: 10.1007/s00438-003-0895-2. [DOI] [PubMed] [Google Scholar]
- 43.Ivanova C, Bååth JA, Seiboth B, et al. Systems analysis of lactose metabolism in Trichoderma reesei identifies a lactose permease that is essential for cellulase induction. PLoS ONE. 2013;8:e62631. doi: 10.1371/journal.pone.0062631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhang W, Kou Y, Xu J, et al. Two major facilitator superfamily sugar transporters from Trichoderma reesei and their roles in induction of cellulase biosynthesis. J Biol Chem. 2013;288:32861–32872. doi: 10.1074/jbc.M113.505826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Porciuncula JO, Furukawa T, Shida Y, et al. Identification of major facilitator transporters involved in cellulase production during lactose culture of Trichoderma reesei PC-3-7. Biosci Biotechnol Biochem. 2013;77:1014–1022. doi: 10.1271/bbb.120992. [DOI] [PubMed] [Google Scholar]
- 46.Ries L, Pullan ST, Delmas S, et al. Genome-wide transcriptional response of Trichoderma reesei to lignocellulose using RNA sequencing and comparison with Aspergillus niger. BMC Genom. 2013;14:1–12. doi: 10.1186/1471-2164-14-541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Dos Santos Castro L, de Paula RG, et al. Understanding the role of the master regulator XYR1 in Trichoderma reesei by global transcriptional analysis. Front Microbiol. 2016;7:175. doi: 10.3389/fmicb.2016.00175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhong Y, Yu H, Wang X, et al. Towards a novel efficient T-DNA-based mutagenesis and screening system using green fluorescent protein as a vital reporter in the industrially important fungus Trichoderma reesei. Mol Biol Rep. 2011;38:4145–4151. doi: 10.1007/s11033-010-0534-z. [DOI] [PubMed] [Google Scholar]
- 49.Zhong Y, Wang X, Yu H, et al. Application of T-DNA insertional mutagenesis for improving cellulase production in the filamentous fungus Trichoderma reesei. Bioresour Technol. 2012;110:572–577. doi: 10.1016/j.biortech.2012.01.129. [DOI] [PubMed] [Google Scholar]


