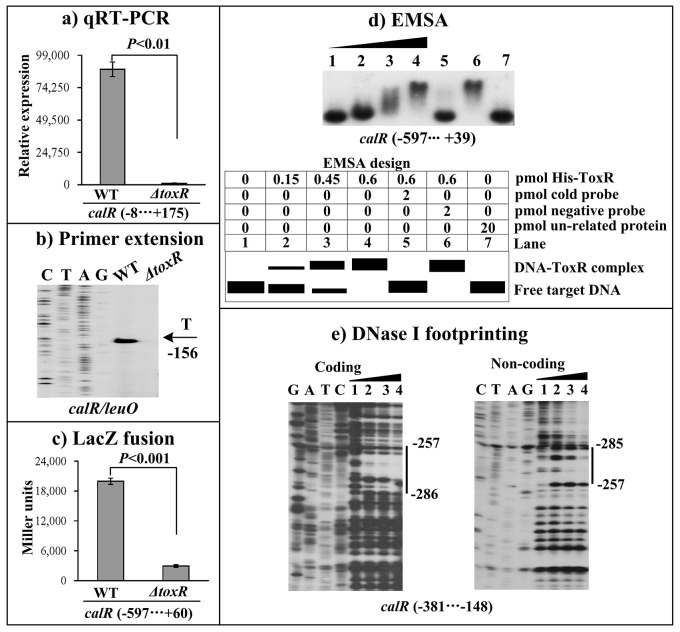
Figure 2. ToxR activates the transcription of calR.
(A) qRT-PCR. The relative mRNA level of calR was compared between ΔtoxR and WT. (B) Primer extension. An oligonucleotide primer was designed to be complementary to the RNA transcript of calR. The primer extension products were analyzed with an 8 M urea -6% acrylamide sequencing gel. The transcription start site is indicated by the arrow with nucleotide and position. (C) LacZ fusion assay. The entire promoter-proximal region of toxR was cloned into pHRP309, and then transformed into WT or ΔtoxR to determine the β-galactosidase activity (miller units) in cellular extracts. (D) EMSA. The entire promoter-proximal region of calR was incubated with increasing amounts of purified His-ToxR protein, and then subjected to 6% (w/v) polyacrylamide gel electrophoresis. Shown below the binding is the schematic representation of the EMSA design. (E) DNase I footprinting. Labeled coding or non-coding DNA probes were incubated with increasing amounts of purified His-ToxR (Lanes 1, 2, 3, and 4 containing 0, 0.2, 0.6, and 0.8 pmol, respectively), and subjected to DNase I footprinting assay. The footprint regions were indicated with vertical bars. The negative and positive numbers represent the nucleotide position upstream and downstream of calR, respectively. Lanes C, T, A and G represent the Sanger sequencing reactions.