Fig. 3.
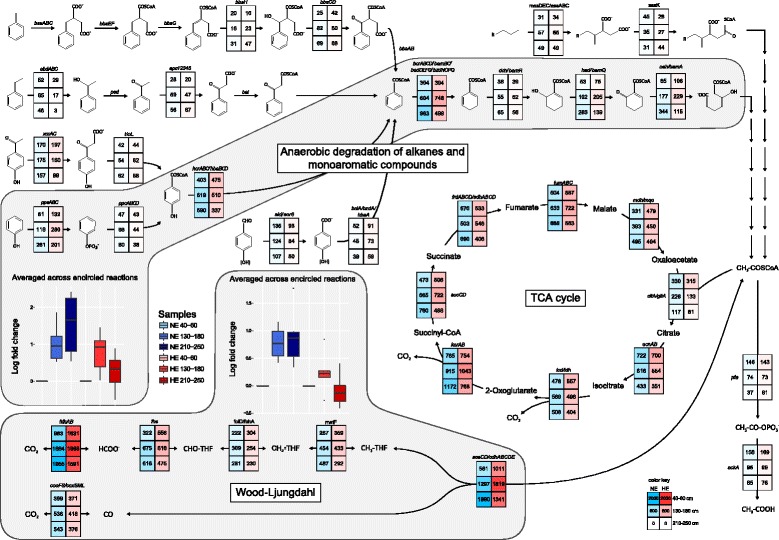
Reconstruction of the complete degradation of n-alkanes and monoaromatic compounds to CO2. The normalized absolute abundances of the genes for each step are given in the respective cells. The selection of the genes for the anaerobic degradation of aromatic compounds was based on the proteogenomics-based reconstruction of their catabolism in the denitrifying Aromatoleum aromaticum EbN1 [78] and the sulfate-reducing Desulfobacula toluolica Tol2 [15]. Only genes with abundances higher than 30 reads in at least one sample are presented. The boxplots depict the log fold changes of the abundances of all genes coding for the enzymes of the anaerobic degradation of phenolic compounds and the Wood-Ljungdahl pathway, respectively. Each sample was compared to the sample at 40–60-cm depth of the same site. Enzyme names: bssABC, benzylsuccinate synthase; bbsEF, succinyl-CoA:(R)-4-isopropylbenzylsuccinate CoA-transferase; bbsG, (R)-benzylsuccinyl-CoA dehydrogenase; bbsH, phenylitaconyl-CoA hydratase; bbsCD, 2-[hydroxy(phenyl)methyl]succinyl-CoA dehydrogenase; bbsAB, benzoylsuccinyl-CoA thiolase; ebdABC, ethylbenzene dehydrogenase; ped, (S)-1-phenylethanol dehydrogenase; apc12345, acetophenone carboxylase; bal., benzoylacetate-CoA ligase; xccAC, 4-hydroxyacetophenone carboxylase; tioL, predicted thiolase; ppsABC, phenylphosphate synthetase; ppcABCD, phenylphosphate carboxylase; hcrAB/hbaBCD, 4-hydroxybenzoyl-CoA reductase; ald/aor6, benzaldehyde dehydrogenase; bclA/bzdA/hbaA, 4-hydroxybenzoate-CoA/benzoate-CoA ligase; bcrABCD/bamBC/badDEFG/bzdNOPQ, benzoyl-CoA reductase; dch/bamR, cyclohex-1,5-diene-1-carbonyl-CoA hydratase; had/bamQ, 6-hydroxycyclohex-1-ene-1-carbonyl-CoA dehydrogenase; oah/bamA, 6-oxocyclohex-1-ene-1-carbonyl-CoA hydrolase; masDEC/assABC, (1-methylalkyl)succinate synthase; assK, AMP-dependent CoA ligase/synthetase; citA/gltA, citrate synthase; acnAB, aconitate hydratase; icd/idh, isocitrate dehydrogenase; korAB, 2-oxoglutarate:ferrodoxin oxidoreductase; sucCD, succinyl-CoA ligase; frdABCD/sdhABCD, fumarate reductase/succinate dehydrogenase; fumABC, fumarate hydratase; mdh/mqo, malate dehydrogenase/malate:quinone oxidoreductase; fdhAB, formate dehydrogenase; fhs, formate-tetrahydrofolate ligase; folD/fchA, methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase; metF, methylenetetrahydrofolate reductase; cooFS/coxSML, carbon monoxide dehydrogenase; acsCD/cdhABCDE, carbon monoxide dehydrogenase/acetyl-CoA synthase; pta, phosphate acetyltransferase; ackA, acetate kinase
