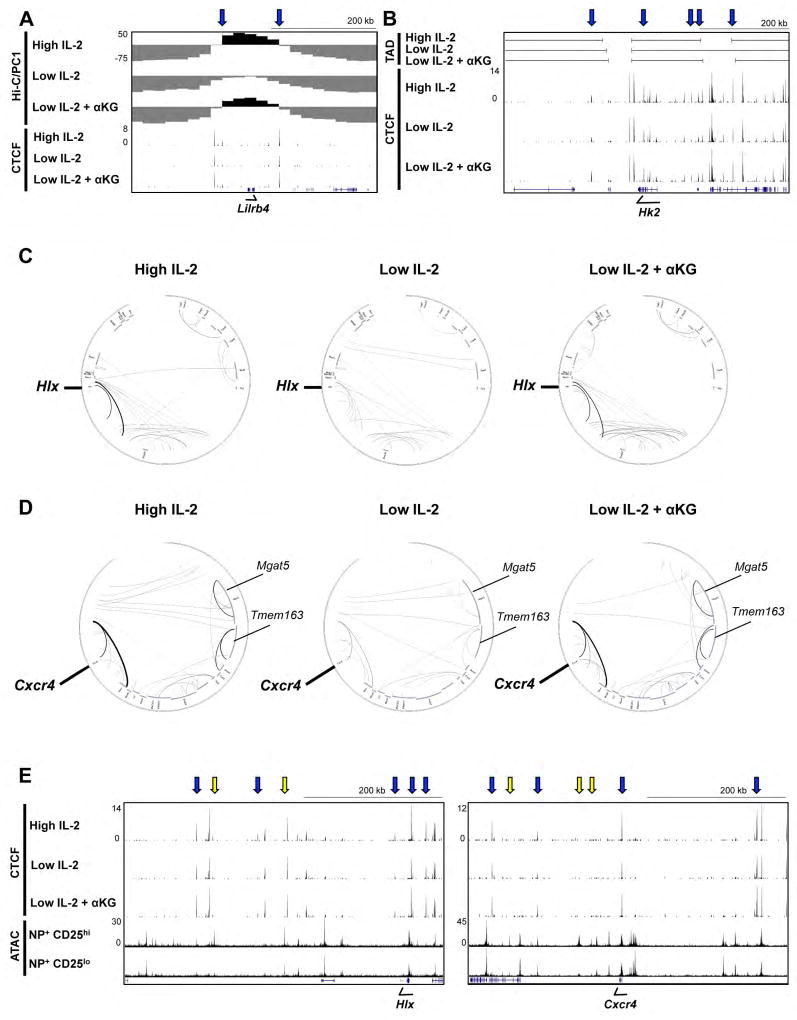
Figure 5. IL-2- and αKG-sensitive CTCF association correlates with changes in genomic organization.
(A–D) In situ Hi-C or CTCF ChIP-seq experiments were performed with primary CD4+ T cells polarized in Th1 conditions and maintained in high IL-2, low IL-2, or low IL-2 with αKG for two days. (A,B) UCSC genome browser tracks displaying a Hi-C (A) PC1 analysis to define transcriptionally permissive (PC1 positive; black) versus inert/or transcriptionally repressive (PC1 negative; grey) genomic compartments, (B) an analysis of outer TAD boundaries and (A,B) ChIP-seq CTCF tracks as described in 4C. (C, D) Circos plots for genomic regions surrounding Hlx (Chr1:184,000,000–187,200,000) and Cxcr4 (Chr1:128,800,000–131,040,000) indicating the probability for genomic interactions from the in situ Hi-C analysis. The minimum probability of interaction shown is a p value of ≤0.0001 with the increased weight of a line indicating a higher significance for the interaction (lower p value). (E) UCSC genome browser tracks displaying CTCF ChIP-seq from Th1 cells exposed to high IL-2, low IL-2, or low IL-2 with αKG, and an ATAC-seq analysis of NP-specific CD8+ T cells isolated 9 days after influenza infection that were sorted into CD25hi or CD25lo populations. Regions displayed were associated with the interaction loops identified in the circos plots in (C, D). Blue arrows indicate IL-2- and αKG-sensitive CTCF sites and yellow arrows indicate ATAC-seq changes. See also Fig. S4–6.