Figure 4.
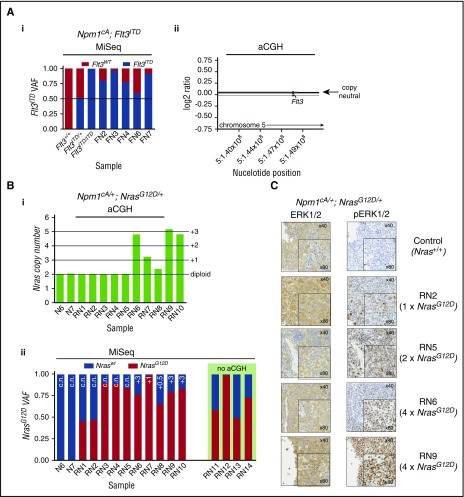
Leukemic progression in double-mutant mice involves increased NrasG12Dor Flt3ITDallele dosage. (A) Increase in Flt3ITD allele burden in AMLs from Npm1cA;Flt3ITD mice through loss of heterozygosity for the locus. (i) Flt3ITD amplicon sequencing (MiSeq) of leukemic bone marrow or spleen DNA (FN2-FN7). Tail DNA amplified from 2-week-old Flt3+/+, Flt3ITD/+, and Flt3ITD/ITD mice were used as control. (ii) Normalized log2 ratio plots show normal copy number for the Flt3 locus in 7 of 7 Npm1cA;Flt3ITD murine AMLs (FN-AMLs) tested. (Bi) Summary of aCGH showing copy number gain at the Nras locus in AMLs RN6-10. (Bii) Allele fractions for Nraswt vs NrasG12D show that copy number gains in RN6-10 involved NrasG12D and that an additional 3 cases (RN3-5) show copy-neutral (c.n.) loss of heterozygosity. In addition, 2 more RN AMLs show gains in mutant NRAS when measuring Nraswt vs NrasG12D allele fractions (aCGH was not performed on these). Results of 2 Npm1cA/+ samples are also shown for comparison purposes (N6, N7). (C) Increased gene dosage of NrasG12D correlates with increased levels of phosphorylated RAS effectors pERK1/2. FN2,3,4,6,7 = NPM1cA/+;Flt3ITD/+ AMLs, RN1-14 = Npm1cA/+;NrasG12D/+ AMLs. VAF, variant allele fraction.