Abstract
Myotonic dystrophy type 1 (DM1) is a multisystemic disease caused by an expanded CTG repeat in the 3′ UTR of the dystrophia myotonica protein kinase (DMPK) gene. Short, DNA-based antisense oligonucleotides termed gapmers are a promising strategy to degrade toxic CUG expanded repeat (CUGexp) RNA. Nucleoside analogs are incorporated to increase gapmer affinity and stability; however, some analogs also exhibit toxicity. In this study, we demonstrate that the 2′,4′-BNANC[NMe] (BNANC) modification is a promising nucleoside analog with high potency similar to 2′,4′-LNA (LNA). BNANC gapmers targeting a nonrepetitive region of the DMPK 3′ UTR show allele-specific knockdown of CUGexp RNA and revert characteristic DM1 molecular defects including mis-splicing and accumulation of RNA foci. Notably, the BNANC gapmers tested in this study did not induce caspase activation, in contrast to a sequence matched LNA gapmer. This study indicates that BNANC gapmers warrant further study as a promising RNA targeting therapeutic.
Graphical abstract

Myotonic dystrophy is a multisystemic disease affecting approximately 1 in 8000 individuals.1 Myotonic dystrophy type I (DM1) is caused by a CTG repeat expansion within the 3′ UTR of the DMPK gene.2,3 Expanded CUG repeat (CUGexp) RNA is retained in the nucleus and forms RNA foci that sequester the MBNL family of splicing factors and induces upregulation of CELF1 through PKC-mediated phosphorylation and altered microRNA regulation.4–6 Altered MBNL and CELF1 activity in DM1 leads to defects in developmentally regulated alternative splicing.1
Antisense oligonucleotides (ASOs) are a promising therapeutic approach for diseases caused by an RNA gain of function, including DM1. The addition of chemical modifications greatly stabilizes the affinity and stability of ASOs. These modifications include altered backbone chemistry, such as phosphorothioate (PS), and altered ribose chemistry, such as the 2′-O-methoxyethyl (MOE) and 2′-O,4′-C-methylenebridged nucleic acid (LNA).7 However, the addition of 2′ ribose modifications abolishes RNase H1-mediated target degradation.8 Chimeric ASOs called gapmers contain 2′ ribose modifications at the 5′ and 3′ ends, leaving an internal 6–10 nucleotide gap that maintains RNase H1 activity.8–10 This is in contrast to mixmers, which contain modified analogs throughout the ASO and are unable to recruit RNase H1 for target cleavage (Figure S1A).8
A recent clinical trial using a MOE gapmer (IONIS-DMPKRx) for the treatment of myotonic dystrophy demonstrated promise with the approach with current work focused on improving tissue delivery (Ionis Pharmaceuticals and Biogen press release, January 2017). Comparison of LNA and MOE gapmers indicates that LNA gapmers are 5–10 fold more potent, eliciting initial interest as a more potent alternative to MOE gapmers.11,12 However, multiple studies have indicated that LNA gapmers induce RNase H1-dependent hepatotoxicity and apoptosis in both mice and cell models, reducing suitability for translational studies.12–15
The 2′4′-BNANC[NMe] (BNANC) modification was developed following similar principles to those of the LNA modification, with a bridged structure that greatly increases its affinity (Figure S1B).16,17 BNANC ASOs are more stable than LNA ASOs and have been shown to be well tolerated in mice.16,18 However, BNANC gapmers have not been systematically tested in repeat expansion models, and it is unknown whether BNANC gapmers have similar toxicity concerns as LNA gapmers. We demonstrate that BNANC gapmers have comparable potency to LNA gapmers, display potentially lower propensity to induce caspase activity, and functionally rescue characteristic DM1 defects, making BNANC gapmers a promising alternative chemistry for therapeutic development. We first sought to compare the efficiency of BNANC and LNA gapmers at targeting DMPK transcripts containing CUGexp. COSM6 cells were transfected with two plasmids: an rtTA expression plasmid and the pBitetDT700ctgGFP (Bi700CTG) plasmid, which contains a bidirectional tet-inducible promoter to express GFP and exons 11–15 of DMPK, including ~300 CUG repeats (Figure 1A). Twenty-four hours after plasmid transfection, cells were treated with increasing concentrations of LNA or BNANC gapmers targeting either within the CUG repeat (CAG gapmers) or in a nonrepetitive region of DMPK (DMPK gapmers; Figure 1; Table S1).19 DMPK mRNA levels were quantified by RT-PCR relative to the GFP mRNA to standardize for transfection efficiency. Compared to the CTG negative control gapmers, both LNA and BNANC CAG gapmers gave significant knockdown of CUGexp RNA at the lowest gapmer dose (0.3 nM) and maximum knockdown to 65–80% of baseline at doses above 10 nM (Figure 1C,D). We next tested the efficiency of LNA and BNANC DMPK gapmers that target upstream of the expanded repeat. While neither DMPK gapmer showed significant knockdown at the lowest dose, they both induced over 70% knockdown at 30 nM. Interestingly, the DMPK gapmers displayed more potent knockdown than the CAG gapmers at the highest dose (100 nM), with only 2–14% of the target remaining (Figure 1C,D). The LNA DMPK gapmer was also more effective than the BNANC DMPK gapmer at the low dosage levels. It is likely that the CAG gapmer is more effective than the DMPK gapmer at low concentrations because there are more binding sites per molecule. At higher concentrations, the residual amount of DMPK RNA that cannot be degraded by the CAG gapmer suggests the CUG repeat is partially protected, potentially by MBNL proteins or RNA structure. Taken together, these results indicate that both LNA and BNANC gapmers are potent and effective for knockdown of CUGexp RNA.
Figure 1.
LNA and BNANC gapmers knockdown of CUGexp RNA. (A) A bidirectional, tetracycline inducible CMV promoter drives expression of both DMPK RNA containing 300 CUG repeats and GFP in the Bi700GFP minigene. RT-PCR primers and gapmer ASO positions are indicated. (B) Transient transfection strategy in COSM6 cells. rtTA = plasmid expressing reverse tetracycline-controlled trans activator. (C) RT-PCR demonstrates DMPK RNA knockdown after treatment with increasing concentrations of LNA and BNANC gapmers. Separate PCR reactions with different cycle numbers were performed for DMPK and GFP using the same cDNA. (D) Quantification of DMPK mRNA knockdown by LNA and BNANC gapmers; n = 3 (CAG gapmers) and n = 4 (DMPK gapmers), Tukey’s multiple comparisons test.
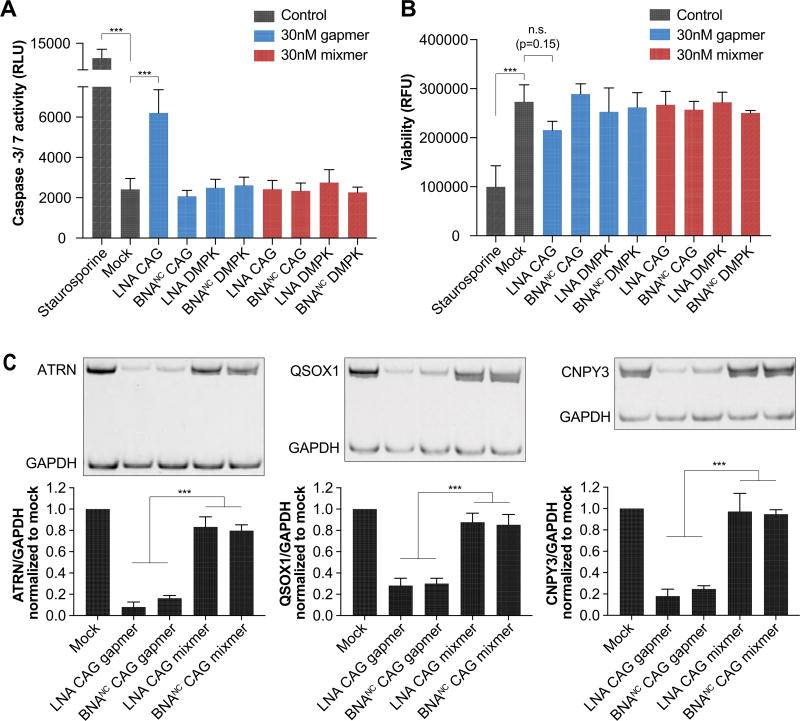
After establishing comparable potency between LNA and BNANC gapmers, we compared the toxicity of BNANC and LNA gapmers. Non-DM1 TeloMyoD fibroblasts were treated with 30 nM of LNA or BNANC gapmers or mixmers and were assayed for membrane integrity, viability, and caspase-3/7 activity 32 h after transfection. We show a sequence-, chemistry-, and RNase H1-dependent increased caspase response specifically caused by the LNA CAG gapmer and not the BNANC CAG gapmer or the LNA CAG mixmer (Figure 2A, Figure S2A). Increased caspase was not due to differences in cell number between samples because only the staurosporine treated cells showed a statistically significant decrease in viability (Figure 2B). Cytotoxicity was only elevated in cells treated with the positive control digitonin, indicating the ASOs tested do not display cytotoxic effects at 30 nM (Figure S2B).
Figure 2.
LNA but not BNANC CAG gapmers induce caspase activity. (A,B) Non-DM1 TeloMyoD fibroblasts were electroporated with 30 nM of LNA or BNANC gapmers or mixmers and plated into differentiation media. Caspase activity (A) and viability (B) were measured 32 h later; n = 3, Dunnett’s multiple comparisons test. (C) RT-PCR of on-target but nondesired knockdown of mRNAs containing CUG6+ normalized to GAPDH in non-DM1 cells; n = 3, Tukey’s multiple comparisons test.
Given that the LNA CAG mixmer did not elicit caspase activity, we hypothesized that LNA CAG gapmers induce caspase activity through on-target but nondesirable knockdown of non-DMPK transcripts that contain CUG6+. However, we demonstrate that both LNA and BNANC CAG gapmers induce RNase H1-mediated knockdown of all 11 tested CUG6+- containing mRNAs (Figure 2C, Figure S3). The question remains why LNA CAG gapmers induce caspase activity but the sequence-matched BNANC CAG gapmers do not. Gapmer-mediated RNase H1 activity includes off-target effects within intronic or exonic regions of long pre-mRNA transcripts.14,20 Additionally, a variety of proteins bind ASOs and modulate their activity, and ASO chemistry can either promote or inhibit protein binding.21,22 Therefore, differences in off-target effects or in the ASO-protein interactome could explain the differential caspase induction seen between the LNA and BNANC CAG gapmers.
Previous work indicates that gapmers targeting within the repetitive region of DMPK preferentially degrade the mutant allele, and this region is an attractive target given the higher number of potential binding sites across the repeat tract.23 However, given our results that non-DMPK transcripts containing short CUG repeat tracts were degraded by CAG gapmers, we analyzed endogenous allele-specific RNA expression in DM1 cells after treatment with BNANC gapmers. To distinguish between the expanded and nonexpanded alleles, we utilized a polymorphic site within DMPK and confirmed that RNA transcripts from the expanded allele are retained in the nucleus in these DM1 cells (Figure 3A–C).24 Treatment with 30 nM BNANC DMPK gapmers induces preferential knockdown of the nuclear-retained CUGexp RNA without affecting the nuclear or cytoplasmic expression of the nonexpanded allele (Figure 3D,E). Unexpectedly, this preferential knockdown was not seen with the BNANC CAG gapmer, which produced over 60% knockdown of total DMPK, but at the expense of targeting both the expanded and nonexpanded alleles (Figure 3E). In contrast, the BNANC DMPK gapmer produced only 20% knockdown of total DMPK, but this was largely accounted for by knockdown of the nuclear-retained, expanded allele. It is possible that the high potency of the BNANC gapmer saturated knockdown of the expanded allele, leading to secondary degradation of the nonexpanded DMPK mRNA. A recent study demonstrated that DMPK knockout mice do not have muscle or cardiac dysfunctions, making it possible that the CAG gapmer would still be viable for clinical translation.25 However, the relative low specificity of this gapmer and higher toxicity seen for the LNA gapmer of this sequence indicate that targeting this repetitive region confers higher risk.
Figure 3.
Allele-specific analysis of DMPK expression in DM1 and non-DM1 TeloMyoD fibroblasts before and after treatment with BNANC gapmers. (A) Diagram of the RT-PCR product containing the BpmI polymorphism. (B,C) RT-PCR and quantification of DMPK allele 1 and allele 2 mRNA levels in the nucleus and cytoplasm from DM1 and non-DM1 TeloMyoD fibroblasts. A control PCR product containing the BpmI site was added to BpmI digestions in lanes 5–8 to confirm complete BpmI digestion. Efficiency of nuclear and cytoplasmic separation is indicated by RTPCR for NEAT1 and S14, respectively. N = nuclear, C = cytoplasmic; n = 3, Sidak’s multiple comparisons test. (D,E) Allele-specific RT-PCR and quantification of DMPK RNA from DM1 fibroblasts after treatment with 30 nM BNANC gapmers. Expression of each allele in either the nucleus or cytoplasm was normalized to GAPDH and displayed relative to the total DMPK expression in mock treated cells. Statistically significant differences in total DMPK mRNA relative to mock are shown for the comparison of CTG control gapmers to CAG and DMPK gapmers; n = 3, Dunnett’s multiple comparisons test.
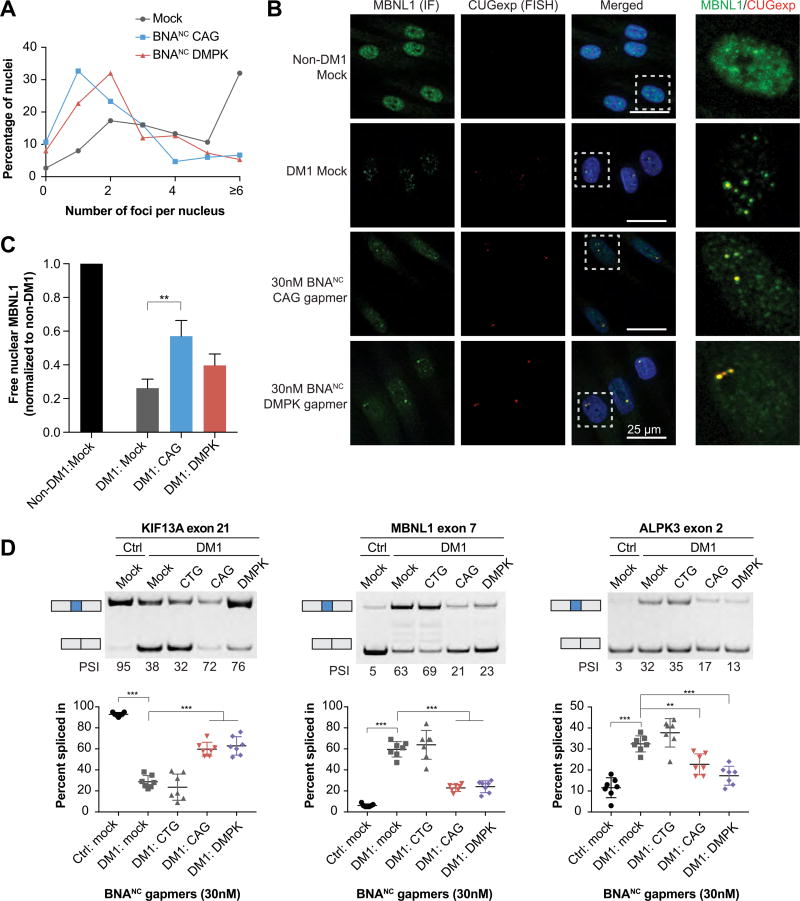
We next sought to determine whether BNANC gapmers could functionally rescue characteristic defects in DM1 cells. First, we used fluorescent in situ hybridization (FISH) and immunofluorescence (IF)-FISH to determine whether BNANC gapmers can reduce foci load and release sequestered MBNL1. DM1 and non-DM1 TeloMyoD fibroblasts were treated with 30 nM of BNANC CAG or DMPK gapmers and differentiated for 6 days to induce the myogenic program.26,27 CUGexp RNA foci were visualized by RNA FISH (Figure S4A), and the number of foci per nucleus was quantified across three biological replicates (Figure 4A). Nearly all (97% ± 2.3%) mock treated DM1 cells had nuclear RNA foci (Figure 4A, Figure S4C), while none of the 150 analyzed non-DM1 cells had nuclear foci. The total number of foci in 150 cells decreased from 738 in DM1 mock treated cells, to 334 and 383 after treatment with BNANC CAG or DMPK gapmers, respectively. This is reflected as a shift in the distribution of foci number per cell (Figure 4A). While over 30% of mock treated cells had more than six foci, only 5–7% of cells treated with the BNANC CAG or DMPK gapmers had more than six foci. This effect was even more pronounced in undifferentiated cells assayed 24 h after gapmer treatment (Figure S4B–D). Partial foci knockdown, despite significant transcript knockdown by RT-PCR, may be a result of foci nucleation by a low absolute number of transcripts.28 It is also possible that the MBNL-bound CUGexp RNA region is partially protected from exonuclease activity following RNase H1 cleavage.
Figure 4.
Rescue of characteristic DM1 defects by BNANC gapmers. (A) The distribution of foci number per nucleus from 150 DM1 TeloMyoD differentiated fibroblasts after treatment with 30 nM BNANC gapmers. (B,C) Quantification and visualization of diffuse nuclear MBNL1 (60 cells per replicate); n = 3, Dunnett’s multiple comparisons test. (D) RT-PCR gels and quantification of splicing rescue for three representative splicing events. Up to two additional PCR cycles were used for ALPK3 in non-DM1 cells compared to DM1 cells, to account for variance in baseline expression levels. PSI = percent spliced in n = 7, Dunnett’s multiple comparisons test.
Given that the majority of cells still had at least one RNA focus after gapmer treatment, we were interested to determine the degree to which MBNL1 was released in cells that had decreased levels of RNA foci. We demonstrated that diffuse MBNL1 levels increased in the nucleoplasm for cells treated with BNANC CAG gapmers; however, this result was not significant in the BNANC DMPK gapmers (Figure 4B,C and Figure S5). The increase in free MBNL1 is visible even in cells that contain foci, indicating that the presence of RNA foci is not necessarily a direct measure of MBNL1 availability. Our results in both differentiated and undifferentiated DM1 cells indicate that RNA foci are reduced, and MBNL1 activity may be increased after treatment with 30 nM of BNANC gapmers.
As RNA foci can be indicative of MBNL1 sequestration in DM1, we next analyzed whether characteristic DM1 splicing abnormalities are rescued by BNANC gapmers. Differentiated DM1 TeloMyoD fibroblasts recapitulate splicing defects observed in DM1 skeletal muscle. Treatment of these cells with 30 nM of BNANC CAG or DMPK gapmers significantly reverted splicing of ALPK3, MBNL1, and KIF13A toward the pattern in non-DM1 cells (Figure 4D). The splicing rescue for MBNL1 and KIF13A was robust, while the rescue was more moderate for the less severely affected event ALPK3. The level of splicing rescue for these events is notable, especially given the residual number of about 1–3 foci per cell after gapmer treatment. This suggests that decreased foci load can be sufficient to revert splicing without complete degradation of all RNA foci and demonstrates that BNANC gapmers potently revert splicing defects in DM1 cells.
In conclusion, we have shown that BNANC gapmers are a promising therapeutic option to target toxic RNA in myotonic dystrophy. The allele-specific knockdown achieved with the BNANC DMPK, but not CAG, gapmers indicates that targeting a nonrepetitive region improves preferential knockdown of the expanded allele. Determining the mechanism responsible for differences in caspase induction between LNA and BNANC CAG gapmers is a key area of interest and will aid in overcoming toxicity concerns for future gapmer-based therapies.
METHODS
Antisense Oligonucleotides
LNA and BNANC ASOs were purchased from Exiqon and Bio-Synthesis, respectively.
Cell Models and Assays
SB TeloMyoD (control, non-DM1) and KB TeloMyoD (DM1, 400 CTG repeats) immortalized human fibroblast cell lines express telomerase and contain a tetracycline-inducible MyoD to promote the myogenic program in response to growth for 6 days in low serum media (1% FBS) supplemented with 1 µg/mL doxycycline.26,27 Toxicity analyses were conducted using the ApoTox-Glo Triplex Assay (Promega). Additional assay details and RT-PCR primers are listed in the Supporting Information and Table S2.
Microscopy
TeloMyoD fibroblasts were electroporated with 30 nM of gapmers, differentiated for 6 days, and visualized by RNA FISH using the (CAG)5 TYE 563 LNA probe (CAGCAGCAGCAGCAG) or by IF/FISH using MBNL1 (Santa Cruz, sc-47740). For further details, see the Supporting Information.
Statistical Analysis
Data are expressed as mean ± standard deviation. All data shown are the summary of three or more biological replicates, and statistical analyses were completed in Prism 7. For data sets where three or more groups were analyzed simultaneously, one-way ANOVA was used with ungrouped data, and two-way ANOVA was used with grouped data, and they were corrected for multiple comparisons. Statistical values used: *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary Material
Acknowledgments
Funding
This work was funded by a National Science Foundation Graduate Research Fellowship [DGE1255980 to K.S.M.]; the National Institutes of Health [5T32 GM008231 to K.S.M.; R01AR045653, R01HL045565, and R01AR060733 to T.A.C.]; the Muscular Dystrophy Association [RG 276796 to T.A.C.]; and Baylor Research Advocates for Student Scientists. This work was also supported by the Integrated Microscopy Core at Baylor College of Medicine, with funding from the National Institutes of Health [DK56338, and CA125123], the Dan L. Duncan Cancer Center, and the John S. Dunn Gulf Coast Consortium for Chemical Genomics.
We are grateful to J. Dubrulle at the Integrated Microscopy Core at Baylor College of Medicine for providing computational image analysis. We thank D. Brook for the TeloMyoD cell lines and P. Sarkar for the expanded pure CTG repeats.
Footnotes
ASSOCIATED CONTENT
The authors declare the following competing financial interest(s): None for T.A.C., K.S.M., and A.N.R. M.C. is the CEO of Bio-Synthesis Inc.
References
- 1.Thornton CA. Myotonic dystrophy. Neurol Clin. 2014;32:705–719. viii. doi: 10.1016/j.ncl.2014.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brook JD, McCurrach ME, Harley HG, Buckler AJ, Church D, Aburatani H, Hunter K, Stanton VP, Thirion JP, Hudson T. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell. 1992;69:385. doi: 10.1016/0092-8674(92)90418-c. [DOI] [PubMed] [Google Scholar]
- 3.Mahadevan M, Tsilfidis C, Sabourin L, Shutler G, Amemiya C, Jansen G, Neville C, Narang M, Barceló J, O’Hoy K. Myotonic dystrophy mutation: an unstable CTG repeat in the 3′ untranslated region of the gene. Science. 1992;255:1253–1255. doi: 10.1126/science.1546325. [DOI] [PubMed] [Google Scholar]
- 4.Jiang H, Mankodi A, Swanson MS, Moxley RT, Thornton CA. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum. Mol. Genet. 2004;13:3079–3088. doi: 10.1093/hmg/ddh327. [DOI] [PubMed] [Google Scholar]
- 5.Kuyumcu-Martinez NM, Wang GS, Cooper TA. Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol. Cell. 2007;28:68–78. doi: 10.1016/j.molcel.2007.07.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kalsotra A, Singh RK, Gurha P, Ward AJ, Creighton CJ, Cooper TA. The Mef2 transcription network is disrupted in myotonic dystrophy heart tissue, dramatically altering miRNA and mRNA expression. Cell Rep. 2014;6:336–345. doi: 10.1016/j.celrep.2013.12.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Prakash TP. An overview of sugar-modified oligonucleotides for antisense therapeutics. Chem. Biodiversity. 2011;8:1616–1641. doi: 10.1002/cbdv.201100081. [DOI] [PubMed] [Google Scholar]
- 8.Kurreck J, Wyszko E, Gillen C, Erdmann VA. Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 2002;30:1911–1918. doi: 10.1093/nar/30.9.1911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Monia BP, Lesnik EA, Gonzalez C, Lima WF, McGee D, Guinosso CJ, Kawasaki AM, Cook PD, Freier SM. Evaluation of 2′-modified oligonucleotides containing 2′-deoxy gaps as antisense inhibitors of gene expression. J. Biol. Chem. 1993;268:14514–14522. [PubMed] [Google Scholar]
- 10.Wu H, Lima WF, Zhang H, Fan A, Sun H, Crooke ST. Determination of the role of the human RNase H1 in the pharmacology of DNA-like antisense drugs. J. Biol. Chem. 2004;279:17181–17189. doi: 10.1074/jbc.M311683200. [DOI] [PubMed] [Google Scholar]
- 11.Jepsen JS, Sørensen MD, Wengel J. Locked nucleic acid: a potent nucleic acid analog in therapeutics and biotechnology. Oligonucleotides. 2004;14:130–146. doi: 10.1089/1545457041526317. [DOI] [PubMed] [Google Scholar]
- 12.Swayze EE, Siwkowski AM, Wancewicz EV, Migawa MT, Wyrzykiewicz TK, Hung G, Monia BP, Bennett CF. Antisense oligonucleotides containing locked nucleic acid improve potency but cause significant hepatotoxicity in animals. Nucleic Acids Res. 2007;35:687–700. doi: 10.1093/nar/gkl1071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kasuya T, Hori S, Watanabe A, Nakajima M, Gahara Y, Rokushima M, Yanagimoto T, Kugimiya A. Ribonuclease H1-dependent hepatotoxicity caused by locked nucleic acid-modified gapmer antisense oligonucleotides. Sci. Rep. 2016;6:30377. doi: 10.1038/srep30377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Burel SA, Hart CE, Cauntay P, Hsiao J, Machemer T, Katz M, Watt A, Bui HH, Younis H, Sabripour M, Freier SM, Hung G, Dan A, Prakash TP, Seth PP, Swayze EE, Bennett CF, Crooke ST, Henry SP. Hepatotoxicity of high affinity gapmer antisense oligonucleotides is mediated by RNase H1 dependent promiscuous reduction of very long pre-mRNA transcripts. Nucleic Acids Res. 2016;44:2093–2109. doi: 10.1093/nar/gkv1210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kakiuchi-Kiyota S, Koza-Taylor PH, Mantena SR, Nelms LF, Enayetallah AE, Hollingshead BD, Burdick AD, Reed LA, Warneke JA, Whiteley LO, Ryan AM, Mathialagan N. Comparison of hepatic transcription profiles of locked ribonucleic acid antisense oligonucleotides: evidence of distinct pathways contributing to non-target mediated toxicity in mice. Toxicol. Sci. 2014;138:234–248. doi: 10.1093/toxsci/kft278. [DOI] [PubMed] [Google Scholar]
- 16.Abdur Rahman SM, Seki S, Obika S, Yoshikawa H, Miyashita K, Imanishi T. Design, synthesis, and properties of 2′,4′-BNA(NC): a bridged nucleic acid analogue. J. Am. Chem. Soc. 2008;130:4886–4896. doi: 10.1021/ja710342q. [DOI] [PubMed] [Google Scholar]
- 17.Miyashita K, Rahman SM, Seki S, Obika S, Imanishi T. N-Methyl substituted 2′,4′- BNANC: a highly nuclease-resistant nucleic acid analogue with high-affinity RNA selective hybridization. Chem. Commun. (Cambridge, U. K.) 2007:3765–3767. doi: 10.1039/b707352f. [DOI] [PubMed] [Google Scholar]
- 18.Yamamoto T, Harada-Shiba M, Nakatani M, Wada S, Yasuhara H, Narukawa K, Sasaki K, Shibata MA, Torigoe H, Yamaoka T, Imanishi T, Obika S. Cholesterol-lowering Action of BNA-based Antisense Oligonucleotides Targeting PCSK9 in Atherogenic Diet-induced Hypercholesterolemic Mice. Mol. Ther.– Nucleic Acids. 2012;1:e22. doi: 10.1038/mtna.2012.16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wheeler TM, Leger AJ, Pandey SK, MacLeod AR, Nakamori M, Cheng SH, Wentworth BM, Bennett CF, Thornton CA. Targeting nuclear RNA for in vivo correction of myotonic dystrophy. Nature. 2012;488:111–115. doi: 10.1038/nature11362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kamola PJ, Kitson JD, Turner G, Maratou K, Eriksson S, Panjwani A, Warnock LC, Douillard Guilloux GA, Moores K, Koppe EL, Wixted WE, Wilson PA, Gooderham NJ, Gant TW, Clark KL, Hughes SA, Edbrooke MR, Parry JD. In silico and in vitro evaluation of exonic and intronic off-target effects form a critical element of therapeutic ASO gapmer optimization. Nucleic Acids Res. 2015;43:8638–8650. doi: 10.1093/nar/gkv857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Liang XH, Sun H, Shen W, Crooke ST. Identification and characterization of intracellular proteins that bind oligonucleotides with phosphorothioate linkages. Nucleic Acids Res. 2015;43:2927–2945. doi: 10.1093/nar/gkv143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liang XH, Shen W, Sun H, Kinberger GA, Prakash TP, Nichols JG, Crooke ST. Hsp90 protein interacts with phosphorothioate oligonucleotides containing hydrophobic 2′-modifications and enhances antisense activity. Nucleic Acids Res. 2016;44:3892–3907. doi: 10.1093/nar/gkw144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lee JE, Bennett CF, Cooper TA. RNase H-mediated degradation of toxic RNA in myotonic dystrophy type 1. Proc. Natl. Acad. Sci. U. S. A. 2012;109:4221–4226. doi: 10.1073/pnas.1117019109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hamshere MG, Newman EE, Alwazzan M, Athwal BS, Brook JD. Transcriptional abnormality in myotonic dystrophy affects DMPK but not neighboring genes. Proc. Natl. Acad. Sci. U. S. A. 1997;94:7394–7399. doi: 10.1073/pnas.94.14.7394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Carrell ST, Carrell EM, Auerbach D, Pandey SK, Bennett CF, Dirksen RT, Thornton CA. Dmpk gene deletion or antisense knockdown does not compromise cardiac or skeletal muscle function in mice. Hum. Mol. Genet. 2016;25:4328–4338. doi: 10.1093/hmg/ddw266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ketley A, Chen CZ, Li X, Arya S, Robinson TE, Granados-Riveron J, Udosen I, Morris GE, Holt I, Furling D, Chaouch S, Haworth B, Southall N, Shinn P, Zheng W, Austin CP, Hayes CJ, Brook JD. High-content screening identifies small molecules that remove nuclear foci, affect MBNL distribution and CELF1 protein levels via a PKC-independent pathway in myotonic dystrophy cell lines. Hum. Mol. Genet. 2014;23:1551–1562. doi: 10.1093/hmg/ddt542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chaouch S, Mouly V, Goyenvalle A, Vulin A, Mamchaoui K, Negroni E, Di Santo J, Butler-Browne G, Torrente Y, Garcia L, Furling D. Immortalized skin fibroblasts expressing conditional MyoD as a renewable and reliable source of converted human muscle cells to assess therapeutic strategies for muscular dystrophies: validation of an exon-skipping approach to restore dystrophin in Duchenne muscular dystrophy cells. Hum. Gene Ther. 2009;20:784–790. doi: 10.1089/hum.2008.163. [DOI] [PubMed] [Google Scholar]
- 28.Gudde AE, González-Barriga A, van den Broek WJ, Wieringa B, Wansink DG. A low absolute number of expanded transcripts is involved in myotonic dystrophy type 1 manifestation in muscle. Hum. Mol. Genet. 2016;25:1648–1662. doi: 10.1093/hmg/ddw042. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.