Abstract
Endothelial cells (EC) act as leading actors in angiogenesis. Understanding the complex network of signal transduction pathways which regulate angiogenesis might offer insights in the regulation of normal and pathological events, including tumours, vascular, inflammatory and immune diseases. The effects of olive oil and of Blueberry extracts upon the phosphoinositide (PI)-specific phospholipase C (PLC) enzymes were evaluated both in quiescent and inflammatory stimulated human umbilical vein EC (HUVEC) using molecular biology (multiliquid bioanalysis) and immunofluorescence techniques. Oleuropein significantly increased the number of surviving HUVEC compared to untreated controls, suggesting that it favours the survival and proliferation of EC. Our results suggest that Oleuropein might be useful to induce EC proliferation, an important event during angiogenesis, with special regard to wound healing. Blueberry extracts increased the number of surviving HUVEC, although the comparison to untreated controls did not result statistically significant. Lipopolysaccharide (LPS) administration significantly reduced the number of live HUVEC. LPS can also modify the expression of selected PLC genes. Adding Blueberry extracts to LPS treated HUVEC cultures did not significantly modify the variations of PLC expression induced by LPS. Oleuropein increased or reduced the expression of PLC genes, and statistically significant results were identified for selected PLC isoforms. Oleuropein also modified the effects of LPS upon PLC genes’ expression. Thus, our results corroborate the hypothesis that Oleuropein owns anti-inflammatory activity. The intracellular localization of PLC enzymes was modified by the different treatments we used. Podosome-like structures were observed in differently LPS treated HUVEC.
Keywords: HUVEC, Endothelial cells, LPS, Phosphoinositide, Plc, Oleuropein, Blueberry, Anthocyanins, Inflammation, Immunofluorescence, Podosome
Introduction
During physiological and pathological events, endothelial cells (EC) act as leading actors in angiogenesis, a complex process in which new vessels arise from the existing vasculature. During the angiogenesis process, required for development, reproduction, wound healing, and tissue regeneration, EC are activated through a multistep cascade (Carmeliet 2003). The morphology, gene expression and activities of EC greatly vary depending on the neighbouring environment (Christ and Wehling 1998, Grienddling and Alexander 1996, Gloe and Pohl 2002). In fact, EC can react to different stimuli with finely, sequentially tuned responses following the recruitment of different signal transduction pathways. That contributes to the adaptation of the endothelium to different situations and needs (Christ and Wehling 1998).
To understand the complex network of signal transduction pathways which regulates angiogenesis might highlight the modulation of normal and pathological events, including tumours, as well as vascular, inflammatory and immune diseases (Ferrara and Davis-Smyth 1997). That might offer a powerful tool for novel therapeutic strategies. The phosphoinositide (PI) specific phospholipase C (PLC) enzymes constitute a complex family belonging to the PI signal transduction pathway. The PI system is involved in many cell activities, including hormone and neurotransmitter signalling, cell growth, membrane and cytoskeleton dynamics, ion channels activity, cell cycle, apoptosis, embryo development and organogenesis, and cell polarity (Bunney and Katan 2011, Comer and Parent 2007, Suh et al. 2008, Nakamura and Fukami 2009).
A combination of compartmentalized and temporal changes in the levels of molecules belonging to the PI pathway, including the phosphatydil inositol bisphosphate (PIP2), can induce a variety of cellular responses. As an example, PIP2 regulates cytoskeleton and membrane dynamics, cytokinesis, nuclear events and activity of selected ion channels (Suh et al. 2008).
PLC enzymes contribute to regulate the spatio-temporal balance of PI molecules (Berridge and Dupont 1994, Divecha and Irvine 1995, Hisatsune et al. 2005), cleaving PIP2 into inositol trisphosphate (IP3) and diacylglycerol (DAG). IP3 induces the release of calcium, a crucial mediator in many cell/tissue activities, such as the progression through the cell cycle, actin polymerisation and related cell migration (Berridge and Dupont 1994; Divecha and Irvine 1995; Hisatsune et al. 2005; Suh et al. 2008; Elsner et al. 1996; Revest et al. 1991). DAG can be cleaved to release further molecules acting in inflammation, such as the arachidonic acid (AA), or can activate the protein kinases C (PKCs). PLC enzymes can be activated by a number of different stimuli, which contributed to divide 13 mammalian isoenzymes into 6 sub-families (Berridge and Dupont 1994; Suh et al. 2008). Beside the mechanisms of regulation, PLC enzymes belonging to the same sub-family also share similarity in aminoacids sequence, and domain organisation. Each PLC enzyme is codified by its own gene, and alternative splicing variants were reported for many isoenzymes (Suh et al. 2008).
The distribution of the PLC enzymes is tissue specific (Suh et al. 2008). The panel of expression of PLCs was demonstrated to change in pathological tissues with respect to the normal counterpart (Lo Vasco et al. 2012a, b; Lo Vasco 2011a, b, Lo Vasco et al., 2010a, b, Lo Vasco et al. 2007, Lo Vasco et al. 2013a, b, c, Di Raimo et al. 2016).
Probably, beside the common role in the cleavage of PIP2, each PLC enzyme bears its own functions in the modulation of physiological and, probably, pathological responses. That might depend on the specific mechanisms of stimulation. These potentialities are far to be highlighted.
In our previous studies in quiescent Human Umbilical Vein EC (HUVEC) we identified a number of PLC enzymes (Lo Vasco et al. 2011b), although inconstantly expressed (Lo Vasco et al. 2013a, b, c). The expression and subcellular localization of PLC enzymes varied after cells underwent different stimuli, such as lipopolysaccharide (LPS) (Lo Vasco et al. 2013a, b, c), fibroblast growth factor (FGF) (Lo Vasco et al. 2014a) or neuropeptide Y (NPY) (Lo Vasco et al. 2014b). Thus, our previous studies suggested that PLC enzymes were involved in the activation of EC under different conditions, such as inflammation, and cell growth stimulation.
In the present paper, we evaluated both in quiescent and inflammatory stimulated HUVEC the effects upon the cell survival and expression of PLC enzymes of olive oil, a traditional Mediterranean diet component, and of Blueberry extracts.
Oleuropein, the main polyphenol in olive oil, was suggested to play a role as protective molecule acting in cell sufferance and death in ageing, as well as in different human diseases, including neurodegenerative or cardiovascular pathologies, cancer and diabetes (Del Rio et al. 2013, Rodrigo et al. 2014, Stefani and Rigacci 2014, Alarcon de la Lastra et al. 2001).
Blueberry constituents are likely to counteract oxidative stress, decrease inflammation, and modulate both interactions among different molecules and gene expression (Neto 2007; Slatnar et al. 2012; Garzón et al. 2010; Duymuş et al. 2014). Blueberries belong to Ericaceae family of berries and represent an important dietary source of bioactive compounds (BAC) bearing antioxidant properties. The bioactive compounds in berries contain phenolic compounds (phenolic acids, flavonoids, such as anthocyanins and flavonols, and tannins) responsible for prevention of inflammation disorders and cardiovascular diseases (Giampieri et al. 2014a; Giampieri et al. 2014b). The bioactive compounds also showed protective effects to lower the risk of various cancers (Seeram et al. 2006, Wedge et al. 2001).
Materials and methods
The following experiments were repeated at least three times.
Cell culture
HUVEC (Cambrex Corporation, Walkersville, MD –USA were cultured as previously described (Lo Vasco et al. 2011). Endothelial Cell Growth Supplement was added to Endogro medium (Endogro, Millipore, MA -USA). Confluent monolayer (>60%) was obtained after 6–9 days. Control cells were maintained for 24 h. Treated cells were stimulated by adding to the medium of culture respectively: i) 100 ng/ml LPS (Sigma-Aldrich, St. Louis, MO); ii) Oleuropein 50 μM (Sigma-Aldrich); iii) Blueberry lyophilized extract 40 μg/ml (see above for preparation of the extract); iv) both LPS and Oleuropein (same dosages previously indicated); v) both LPS and Blueberry extract (same dosages previously indicated). Both treated cells and untreated negative controls, previously plated on six-well plates, were then suspended in TRIzol reagent (Invitrogen Corporation, Carlsbad, CA) and checked after 3, 6, and 24 h.
The lyophilized powder of Blueberry was prepared as previously described (Li et al. 2013; Cesa et al. 2017). Briefly, fresh and fully ripe Blueberry fruits coming from the Trentino region of Italy were washed, dried on paper and homogenized at room temperature for 2 min either by a domestic mixer at 16000 rpm or by a T18 Ultra-Turrax® homogenizer (IKA®, Staufen, Germany) at 10000 rpm. As previously described (Li et al. 2013), the resulting fruit puree was heated at 70 °C for 2 h and rapidly ice-cooled. Then, the mix of anthocyanins was extracted. The homogenates were stirred with 50 mL of ethanol:0.5% acetic acid (70:30, v/v) (Carlo Erba Milan, Italy) (Fluka, Milan, Italy). The suspension was centrifuged at 12000 g for 10 min at 4 °C. The collected supernatant was concentrated under reduced pressure at 40 °C with a rotary evaporator and added with ethanol:0.5% acetic acid (70:30, v/v) to a final volume of 20 ml. The extraction was checked by HPLC analysis by using Perkin-Elmer apparatus equipped with a series LC 200 pump, a series 200 diode array detector and a series 200 autosampler; data were acquired with Perkin-Elmer Totalchrom software. In each sample, the following anthocyanins were identified: delphinidin-3-O-galactoside (del-3-gal), cyanidin-3-O-galactoside (cya-3-gal), delphinidin-3-O-arabinoside (del-3-ara), petunidin-3-O-galactoside (pet-3-gal), cyanidin-3-O-arabinoside (cya-3-ara), petunidin-3-O-arabinoside (pet-3-ara), malvidin-3-O-galactoside (mal-3-gal), malvidin-3-O-glucoside (mal-3-glu) and malvidin-3-O-arabinoside (mal-3-ara) (Cesa et al. 2017).
MTT test
The cell viability was evaluated using colorimetric microassay (Mosmann 1993). Briefly, tetrazolium salt (MTT: 3-[4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazoliumbromide, 5 mg/ml suspended in PB) (Sigma) was added to cell cultures and then incubated for 4 h. The plates were read with a multiwell scanning Elisa reader (wavelength 540 nm, reference length 630 nm) (Biorad Laboratories, Hercules, CA, USA). The relative absorbance was expressed as a percent of the corresponding control and calibrated against control cell cultures plated at different densities, which were then counted in a Coulter ZM cell counter equipped with Channelyser to control for dead cells. The accuracy of the MTT method was shown to be +/−200 cells as previously described (Mosmann 1993).
RT-PCR and molecular biology experiments (Multiliquid bioanalysis)
The experiments were conducted after 24 h from treatment (T0 = seeding time).
HUVEC, both treated and untreated controls, were plated on six-well plates, and suspended in TRIzol reagent (Invitrogen Corporation, Carlsbad, CA, USA) 3, 6 and 24 h after addiction to the medium of culture respectively of: i) 100 ng/ml LPS; ii) Oleuropein 50 μM; iii) Blueberry lyophilized extract 40 μg/ml; iv) LPS plus Oleuropein; v) LPS plus Blueberry extract. Total RNA was isolated from samples following the manufacturer’s instructions. The obtained RNA purity was assessed by using a UV/visible spectrophotometer (SmartSpec 3000, Biorad Laboratories). Total RNA (1 mg) was reverse transcribed by using High Capacity cDNA Reverse Transcription Kit (Applied Byosystems, Carlsbad, CA) according to manufacturer’s instructions. Briefly, RT Buffer, dNTP Mix, RT Random Primers, Multiscribe Reverse Transcriptase, RNase Inhibitor and DEPC-treated distilled water were added in RNase free tubes on ice. RNA sample was added. Thermal cycler program was: 25 °C for 10 min; 37 °C for 120 min; the reaction was stopped at 85 °C for 5 min. The final volume was 20 ml. For PCR reactions, the primer pairs (Bio Basic Inc., Amherst- NY, USA) used for PLC isoenzymes are listed in Table 1.
Table 1.
List of PLC primers for PCR experiments
| PLCB1 | OMIM *607120 | forward 5′-AGCTCTCAGAACAAGCCTCCAACA-3’ |
| reverse 5′-ATCATCGTCGTCGTCACTTTCCGT-3’ | ||
| PLCB2 | OMIM *604114 | forward 5′-AAGGTGAAGGCCTATCTGAGCCAA-3’ |
| reverse 5′-CTTGGCAAACTTCCCAAAGCGAGT-3’ | ||
| PLCB3 | OMIM *600230 | forward 5′-TATCTTCTTGGACCTGCTGACCGT-3’ |
| reverse 5′-TGTGCCCTCATCTGTAGTTGGCTT-3’ | ||
| PLCB4 | OMIM *600810 | forward 5′-GCACAGCACACAAAGGAATGGTCA-3’ |
| reverse 5′-CGCATTTCCTTGCTTTCCCTGTCA-3’ | ||
| PLCG1 | OMIM *172420 | forward 5′-TCTACCTGGAGGACCCTGTGAA-3’ |
| reverse 5′-CCAGAAAGAGAG CGTGTAGTCG-3’ | ||
| PLCG2 | OMIM *600220 | forward 5′-AGTACATGCAGATGAATCACGC-3’ |
| reverse 5′-ACCTGAATCCTGATTTGACTGC-3’ | ||
| PLCD1 | OMIM *602142 | forward 5′-CTGAGCGTGTGGTTCCAGC-3’ |
| reverse 5′-CAGGCCCTCGGACTGGT-3’ | ||
| PLCD3 | OMIM *608795 | forward 5′-CCAGAACCACTCTCAGCATCCA-3’ |
| reverse 5′-GCCA TTGTTGAGCACGTAGTCAG-3’ | ||
| PLCD4 | OMIM *605939 | forward 5′-AGACACGTCCCAGTCTGGAACC- 3’ |
| reverse 5′-CTGCTTCCTCTTCCTCATATTC- 3 | ||
| PLCE | OMIM *608414 | forward 5′-GGGGCCACGGTCATCCAC-3’ |
| reverse 5′-GGGCCTTCATACCGTCCATCCTC-3’ | ||
| PLCH1 | OMIM *612835 | forward 5′-CTTTGGTTCGGTTCCTTGTGTGG-3’ |
| reverse 5′-GGATGCTTCTGTCAGTCCTTCC-3’ | ||
| PLCH2 | OMIM *612836 | forward 5′-GAAACTGGCCTCCAAACACTGCCCGCCG-3’ |
| reverse 5′-GTCTTGTTGGAGATGCACGTGCCCCTTGC-3’ | ||
| GAPDH | GAPDH | forward 5′-CGAGATCCCTCCAAAATCAA-3’ |
| reverse 5′-GTCTTCTGGGTGGCAGTGAT-3’ | ||
| Vil2 | Vil2 | forward 5′ - GCTGTGCAGGCCAAGTTT-3’ |
| reverse 5′ -TCCCACTGGTCC CTGGTA-3’ |
Glyceraldehyde-3-phosphate dehydrogenase (GAPDH; OMIM *138400) housekeeping gene was used as a positive control. The following primer pairs for GAPDH were used (Bio Basic Inc) forward 5′-CGAGATCCCTCCAAAATCAA-3′ and reverse 5′-GTCTTCTGGGTGGCAGTGAT-3′. The specificity of all primers was verified searching in NCBI data base possible homology to cDNAs of unrelated proteins. Positive control cells (osteosarcoma HS888 cells) were used to check the efficacy of the primers for PLCD4 and PLCE.
Each PCR tube contained the following reagents: 0.2 μM of both sense and antisense primers, 1–3 μl (about 1 μg) template cDNA, 0.2 mM dNTP mix, 2.5 U REDTaq Genomic DNA polymerase (Sigma, Germany) and 1X reaction buffer. MgCl2 was added at variable (empirical determination by setting the experiment) final concentration. The final volume was 50 μl. The amplification was started with an initial denaturation step at 94 °C for 2 min and was followed by 35 cycles consisting of denaturation (30 s) at 94 °C, annealing (30 s) at the appropriate temperature for each primer pair and extension (1 min) at 72 °C. The PCR products were analysed by 1.5% TAE ethidium bromide stained agarose gel electrophoresis (Agarose Gel Unit, Biorad Laboratories Inc., UK). PC-assisted CCD camera UVB lamp (Vilber Lourmaret, France) was used for gel documentation. Gel electrophoresis of the amplification products revealed single DNA bands with nucleotide lengths as expected for each primer pair. Optical densities were normalised to the mRNA content of GAPDH. To exclude possible DNA contamination during the RT-PCR, RNA samples were amplified by PCR without reverse transcription. No band was observed, confirming no DNA contamination in the RNA preparation procedure (data not shown). We measured mRNA concentrations as follows: in untreated HUVEC we measured the basal (t = 0) concentration, and after 3, 6 and 24 h to evaluate the effects of the progression of the cell cycle upon transcription. In HUVEC treated with different molecules (LPS, Oleuropein, Blueberry extract), we measured mRNA concentration after 3, 6 and 24 h. The PCR products were analysed and quantified with the Agilent 2100 bioanalyzer using the DNA 1000 LabChip kit (Agilent Technologies, Germany). All the experiment sets were repeated at least three times.
Immunofluorescence
Immunofluorescence analysis of HUVEC was used to evaluate the sub-cellular distribution of PLC enzymes in control HUVEC, in LPS treated HUVEC, in HUVEC treated with LPS and Blueberry extract and in HUVEC treated with LPS and Oleuropein. Briefly, cells were washed three times with PBS and fixed with 4% paraformaldehyde (PFA) in phosphate buffer saline (PBS) for 10 min at 4 °C, then three washes with PBS followed. Cells were incubated with primary antibodies diluted in PBS for 1 h at room temperature. Cover-slips were then incubated with the specific secondary antibody Texas Red or fluorescein-conjugated for 1 h at room temperature. The primary anti-human PLC antibodies and the appropriated fluorescence conjugated secondary antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). Cells were washed twice with 1X PBS 5 min, then counterstained with 4′,6-diamidino-2-phenylindole (DAPI) fluorescent staining to detect the cell nuclei. The slides (2 well μ-slides) (Ibidi, Germany) were visualized and captured with an Olympus IX50 inverted fluorescence microscope (Olympus, Tokyo, Japan) and processed using Adobe Photoshop 7.0 software.
Results
Effects of treatments upon HUVEC survival (MTT test)
The media of results of all the sets of experiments were listed in Tables 2 and 3. The experiments were repeated three times for each observation period (0, 3, 6 and 24 h) and for each treatment. Mathematical media was calculated.
Table 2.
Survival of cells. Media of the results
| T0 | 3h | 6h | 24h | |
|---|---|---|---|---|
| 192.333 | ||||
| Ctrl | 82.500 | 83.958 | 73.333 | |
| Oleuropein | 142.083 | 162.083 | 193.333 | |
| Blueberry | 120.833 | 121.250 | 196.667 | |
| Lps | 71.667 | 95.417 | 95.833 | |
| Lps + Oleuropein | 122.500 | 140.417 | 159.583 | |
| Lps + Blueberry | 63.750 | 115.833 | 165.000 |
Ctrl = controls (untreated cells). T0 = seeding of cells
Table 3.
Comparison of the survival rates of controls and differently treated HUVEC with ANOVA analysis
| p value | |
|---|---|
| Ctrl vs LPS | 0.423272 |
| Ctrl vs LPS + Blueberry extract | 0.300808 |
| Ctrl vs LPS + Oleuropein | 0.004921 |
| Ctrl vs Blueberry extract | 0.059531 |
| Ctrl vs Oleuropein | 0.004921 |
| Ctrl vs treated HUVEC | 0.0294 |
P value was considered significant when <.05
In control HUVEC the survival rate was 42.9% after 3 h. After 6 h, the surviving cells’ number slightly increased 1.74% (43.65% of the initial controls). After 24 h, the surviving cells’ number further reduced about 12.65% (38.1% of the initial controls) (Fig. 1; Table 2).
Fig. 1.
Histogram of HUVEC survival after 3, 6 and 24 h. T0 = seeding time. On y axis: CTRL = untreated HUVEC. OL = HUVEC treated with Oleuropein. BB = HUVEC treated with Blueberry extract. LPS/OL = HUVEC treated with LPS plus Oleuropein. LPS/BB = HUVEC treated with LPS plus Blueberry extract. On x axis: number of live HUVEC. Standard error bars are indicated
After oil treatment, the survival rate of HUVEC was significantly higher than controls after 3, 6 and 24 h (Fig. 1; Table 2). Comparison of the survival rates of controls and oil treated HUVEC with ANOVA analysis resulted statistically significant with p-value = .004921 (Table 3).
After Blueberry extract treatment, the survival rate of HUVEC was higher than controls after 3 and 6 h and even higher after 24 h (Fig. 1; Table 2). Comparison of the survival rates of controls and HUVEC treated with Blueberry extracts with ANOVA analysis did not result statistically significant with p-value = .059531 (Table 3).
After LPS treatment, the survival rate of HUVEC was markedly reduced after 3 h. The number of cells slightly increased after 6 h and stayed about the same after 24 h (Fig. 1; Table 2). Comparison of the survival rates of controls and LPS treated HUVEC with ANOVA analysis did not result statistically significant with p-value = .423272 (Table 3).
After combined treatment with both LPS and Oleuropein, the number of surviving cells was slightly reduced after 3 h, while it slightly and progressively increased after 6 and 24 h (Fig. 1; Table 2). Comparison of the survival rates of controls and oil treated HUVEC resulted statistically significant with p-value = .005572 (Table 2).
After combined treatment with both LPS and Blueberry extract, the number of surviving cells was markedly reduced, although it progressively increased after 6 and 24 h (Fig. 1). Comparison of the survival rates of controls and oil treated HUVEC with ANOVA analysis did not result statistically significant with p-value = .300808 (Table 3).
The overall analysis of the results with ANOVA analysis of statistical variation resulted statistically significant with a p value = 0.0294 (Table 3).
RT-PCR
The results are listed in Table 4. The multiliquid bioanalysis was conducted after 24 h from each treatment. PLCD4 and PLCE were not expressed both in control and treated HUVEC. In positive control cells (osteosarcoma HS888 cells) both PLCD4 and PLCE were detected, as expected (data not shown).
Table 4.
Media of results of Multiliquid Bioanalysis of PCR products for PLC genes
| PLCB1 | PLCB2 | PLCB3 | PLCB4 | PLCD1 | PLCD3 | PLCG1 | PLCG2 | PLCH1 | PLCH2 | GAPDH | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ctrl | 3.78 | 3.69 | 4.28 | 2.03 | 4.46 | 7.33 | 9.59 | 1.2 | 1.85 | 0.7 | 4.2 |
| Blueberry | 2.58 | 2.85 | 4.99 | 0.92 | 4.84 | 2.45 | 5.37 | 0.9 | 0 | 0.58 | 5.77 |
| Oleuropein | 4.48 | 1.44 | 7.01 | 4.89 | 8.27 | 1.64 | 13.4 | 3.41 | 0 | 0.03 | 4.73 |
| LPS | 3.01 | 0 | 5.73 | 0.65 | 3.95 | 1.56 | 1.09 | 0 | 0 | 0.38 | 5.23 |
| LPS + Blueberry | 0 | 0.44 | 5.75 | 1.43 | 3.84 | 2.33 | 5.12 | 0 | 0 | 0.95 | 4.73 |
| LPS + Oleuropein | 4.77 | 0 | 7.57 | 3.2 | 6.52 | 2.27 | 9.04 | 1.89 | 0 | 0.55 | 5.43 |
Ctrl: control HUVEC. Blueberry: HUVEC treated with Blueberry Extract. Oleuropein: HUVEC treated with Oleuropein. LPS: HUVEC treated with LPS. LPS + Blueberry: HUVEC treated with LPS plus Blueberry extract. LPS + Oleuropein: HUVEC treated with LPS plus Oleuropein
All the treatments slightly increased the expression of GAPDH in a statistically non-significant measure (Tables 3 and 4). All the treatments blocked the expression of PLCH1 (Table 4). As the expression of PLCH1 in HUVEC controls is low, that observation did not result statistically significant (Table 5).
Table 5.
Effects of different treatments upon the expression of PLC genes. P value of rates between controls and differently treated HUVEC
| PLCB1 | PLCB2 | PLCB3 | PLCB4 | PLCD1 | PLCD3 | PLCG1 | PLCG2 | PLCH1 | PLCH2 | GAPDH | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ctrl/LPS | .366552 | .006836 | .230904 | .137551 | .416028 | .028575 | .011501 | .035265 | .148333 | .272629 | .212646 |
| Ctrl/LPS+BB | .045171 | .129659 | .287836 | .355426 | .41734 | .144955 | .241 | .035265 | .148333 | .35068 | .313054 |
| Ctrl/LPS+O | .395257 | .101851 | .011789 | .275052 | .192242 | .142552 | .466665 | .019326 | .148333 | .427823 | .182645 |
| Ctrl/BB | .300695 | .401299 | .378535 | .228635 | .44561 | .150193 | .265179 | .252186 | .148333 | .446458 | .10019 |
| Ctrl/O | .369124 | .217272 | .101051 | .116759 | .027475 | .118864 | .298076 | .242198 | .148333 | .017749 | .318127 |
Numbers in italic bold mark the statistically significant results
Ctrl/ LPS: Control HUVEC versus LPS treated HUVEC. Ctrl/ LPS + BB: Control HUVEC versus HUVEC treated with LPS plus Blueberry Extract. Ctrl/ LPS + O: Control HUVEC versus HUVEC treated with LPS plus Oleuropein. Ctrl/ BB: Control HUVEC versus Blueberry Extract treated HUVEC. Ctrl/ O: Control HUVEC versus Oleuropein treated HUVEC
Different LPS treatments. Administration of LPS to HUVEC cultures blocked the expression of PLCB2 and PLCG2; reduced the expression of PLCB1, PLCB4, PLCD1, PLCD3, PLCG1 PLCH2 and increased the expression of PLCB3 (Table 4). The reduction of expression resulted statistically significant for the following genes: PLCB2, PLCD3, PLCG1 and PLCG2 (Table 4). Adding LPS and Blueberry extracts to the cultures blocked the expression of PLCB1 and PLCG2; reduced the expression of PLCB2, PLCB4, PLCD1, PLCD3 and PLCG1 and increased the expression of PLCB3 and PLCH2 (Table 3). The reduction resulted statistically significant for PLCB1 and PLCG2 (Table 5). Adding LPS and Oleuropein to HUVEC cultures blocked the expression of PLCB2; reduced the expression of PLCD3, PLCG1 and PLCH2 and increased the expression of PLCB1, PLCB3, PLCB4, PLCD1 and PLCG2 (Table 4). The increase resulted statistically significant for PLCB3 and PLCG2.
Blueberry extract. Administration of Blueberry extract to HUVEC cultures reduced the expression of PLCB1, PLCB2, PLCB4, PLCD3, PLCG1, PLCG2 and PLCH2, and increased the expression of PLCB3 and PLCD1 (Table 4). However, the variation of PLC genes’ expression induced by Blueberry extract did not result statistically significant (Table 5).
Oleuropein. Administration of Oleuropein to HUVEC cultures reduced the expression of PLCB2, PLCD3 and PLCH2 and increased the expression of PLCB1, PLCB3, PLCB4, PLCD1, PLCG1 and PLCG2 (Table 4). The reduction of PLCH2 expression and the increase of PLCD1 expression resulted statistically significant.
Immunofluorescence
Control HUVEC cells. PLC β1 was detected in the cytoplasm, with granule distribution. PLC β2 was detected in the nucleus (Fig. 2). PLC β3 was detected in the nucleus and in the cytoplasm nearby the perinuclear area (Fig. 2). PLC β4 was detected in a limited number of cells, partly in the nucleus and cytoplasm, partly in the cytoplasm (Fig. 2). PLC δ4 was detected in the nucleus (Figure 3). PLC ε was detected in the nucleus and enhanced in the nuclear membrane (Fig. 3). PLC γ1 was detected in the cytoplasm with granule distribution (Fig. 4). PLC η1 was detected in the nucleus; it was also detected in the cytoplasm, with granule distribution (Fig. 4). PLC η2 was detected in the perinuclear area (figure 4). PLC δ1, PLC δ3 and PLC γ2 were not detected (Figs. 3 and 4).
Fig. 2.
Immunofluorescence of PLC enzymes of the β subfamily. CTRL = control HUVEC; LPS = LPS treated HUVEC; LPS/BB = HUVEC treated with LPS plus Blueberry extract; LPS/O = HUVEC treated with LPS plus Oleuropein. For description, please see the results section
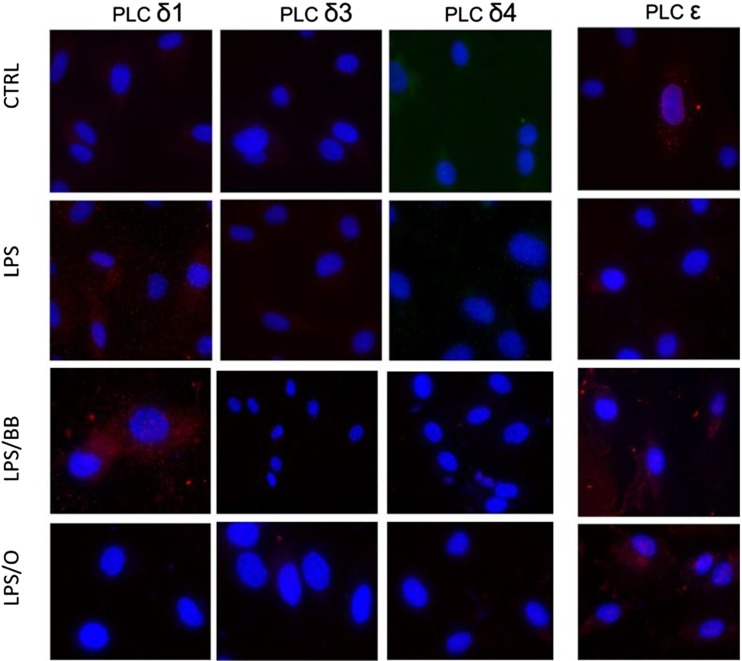
Fig. 3.
Immunofluorescence of PLC enzymes of the δ subfamily and of PLC ε. CTRL = control HUVEC; LPS = LPS treated HUVEC; LPS/BB = HUVEC treated with LPS plus Blueberry extract; LPS/O = HUVEC treated with LPS plus Oleuropein. For description, please see the results section
Fig. 4.
Immunofluorescence of PLC enzymes of the γ and η subfamilies. CTRL = control HUVEC; LPS = LPS treated HUVEC; LPS/BB = HUVEC treated with LPS plus Blueberry extract; LPS/O = HUVEC treated with LPS plus Oleuropein. For description, please see the results section
LPS treated HUVEC. PLC β1 was detected in the cytoplasm, with fine granule distribution, and in podosome-like structures (Fig. 2). PLC β3 was detected in the perinuclear area cytoplasm (Fig. 2). PLC β4 was detected in the nucleus and cytoplasm in a limited number of cells, while it was detected exclusively in the cytoplasm in the remaining cells (Fig. 2). PLC δ3 and PLC γ1 were detected in the nucleus (Figs. 3 and 4). PLC η1 was weakly visible in the nucleus, while it was detected in the perinuclear area (Fig. 4). PLC η2 was detected in the nucleus and in cell evaginations reaching out from cell to cell (Figs. 4 and 5). PLC β2, PLC δ1, PLC δ4, PLC ε and PLC γ2 were not detected (Figs. 2, 3 and 4).
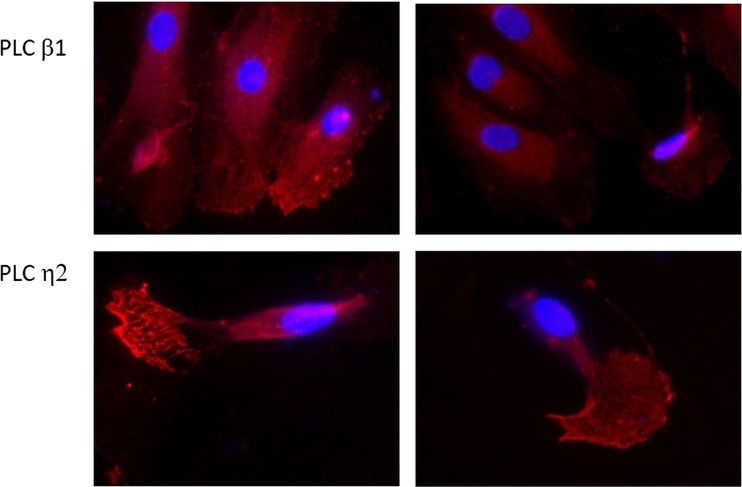
Fig. 5.
Immunofluorescence of PLC β1 (upper images) in LPS treated HUVEC and of PLC η2 (lower images) in HUVEC treated with LPS plus Blueberry extracts. Presence of the enzymes in podosome-like structures
HUVEC treated with both LPS and Blueberry extracts. PLC β1 was detected in the cytoplasm, with granule distribution particularly dense around the nucleus (Fig. 2). PLC β2 was weakly detected in the cytoplasm (Fig. 2). PLC β3 was weakly detected in the cytoplasm, particularly evident in the perinuclear area (Fig. 2). PLC β4 and PLC δ1 were weakly detected in the cytoplasm (Fig. 2 and 3). PLC ε was detected in the nucleus; it was also detected in the perinuclear area of the cytoplasm (Fig. 3). PLC γ1 and PLC γ2 were detected in the nucleus v. PLC η1 was detected in the cytoplasm (Fig. 4). PLC η2 was detected in the nucleus, in the cytoplasm with a submembrane enhancement and in podosome-like structures (Figs. 4 and 5). PLC δ3 and PLC δ4 were not detected (Fig. 3).
HUVEC treated with both LPS and Oleuropein. PLC β1 was detected in the cytoplasm with fine granule distribution (Fig. 2). PLC β2 was detected in the perinuclear area with fine granule distribution (Fig. 2). PLC β3 was detected in the cytoplasm, particularly dense in the perinuclear area (Fig. 2). PLC β4 was weakly detected in the cytoplasm (Fig. 2). PLC ε was detected in the nucleus and weakly in the cytoplasm (Fig. 3). PLC γ1 was detected in the nucleus and in the perinuclear area, where it seems to be particularly concentrated (Fig. 4). PLC γ2 and PLC η2 were detected in the nucleus and in the perinuclear region of the cytoplasm (Fig. 4). PLC η1 was detected in the cytoplasm with granule distribution (Fig. 4). PLC δ1, PLC δ3 and PLC δ4 were not detected (Fig. 3).
Discussion
Our present results indicate that Oleuropein administration to HUVEC cultures increased the number of surviving cells compared to untreated controls. Notably, Oleuropein significantly reduces the cell death rate after seeding (Fig. 1; Table 2). The administration of Blueberry extracts also increased the number of HUVEC, however the results were not statistically significant (Tables 2 and 3).
Confirming our previous works (Lo Vasco et al. 2007; Lo Vasco et al. 2013a, b, c), LPS administration to HUVEC cultures significantly reduced the number of live cells, especially in the 0–3 h interval (Fig. 1, Tables 2 and 3). That suggests how inflammation affects HUVEC survival especially briefly after stimulation. Although the combination of LPS plus Oleuropein treatment reduced the number of surviving HUVEC with respect to untreated controls, the number of live cells was greater compared to LPS treated HUVEC. That might suggest that Oleuropein can modulate the effect of LPS upon cell survival, reducing the number of dead cells. The combination of LPS and Blueberry extract treatment did not significantly modify the effects of LPS treatment alone.
Therefore, Oleuropein seems to favour the survival and proliferation of quiescent HUVEC and to reduce the cell death rate induced by LPS. That confirms the anti-inflammatory role of Oleuropein. Our results suggest that Oleuropein might be useful to induce EC proliferation, an important event during angiogenesis, with special regard to wound healing.
The present results confirm our previous observations (Lo Vasco et al. 2007, Lo Vasco et al., 2010a, b; Lo Vasco et al., 2012a, b; Lo Vasco et al. 2013a, b, c) that LPS can modify the expression of some PLC genes. Adding Blueberry extracts to LPS treated HUVEC cultures did not significantly modify the variations of PLC expression induced by LPS. However, for selected PLC genes, the administration of Blueberry seems to slightly reduce the effects of LPS. In fact, LPS reduced the expression of a number of PLC genes, while adding Blueberry extract to LPS the reduction was less evident (Table 3). That is a surprising observation, in that Blueberry extract alone also reduced the expression of the same genes. However, those results were not statistically significant and further studies are required in order to evaluate the controversial effects of contemporary administration of LPS and Blueberry extract.
All the treatments increased the expression of PLCB3 contemporarily reducing the expression of PLCD3. Although no explanation is currently possible, that might represent an interesting observation requiring further analysis addressed to evaluate the possible relationship between the expression of these two genes.
Oleuropein increased or reduced the expression of PLC genes, and statistically significant results were identified for PLCD1 and PLCH2. Moreover, Oleuropein could modify the effects of LPS upon PLC genes’ expression. The expression of PLCB1, PLCB3, PLCB4, PLCD1 and PLCG1 were respectively reduced by LPS and increased by Oleuropein. Adding contemporarily Oleuropein and LPS to HUVEC cultures attenuated the LPS induced reduction of genes’ expression (Table 3). For PLCG2, this effect was more evident. In fact, the expression of PLCG2 was blocked by LPS, as well as by LPS plus Blueberry extract. In presence of LPS plus Oleuropein, PLCG2 was measured and its expression was increased with respect to untreated control HUVEC. Oleuropein administration markedly increased the expression of PLCG2 (Table 2 and 3).
Other genes, such as PLCB2, PLCD3 and PLCH2 also deserve comments. The expression of those genes was reduced both by LPS and Oleuropein treatment. However, the contemporary administration of LPS and Oleuropein induced a milder reduction of PLCD3 and PLCH2 with respect to controls. On the contrary, the expression of PLCB2 was blocked by LPS treatment and reduced by Oleuropein treatment. The expression of PLCB2 was blocked by the contemporary administration of LPS and Oleuropein, suggesting that Oleuropein did not efficiently counteract the effects of LPS upon this gene.
Therefore, our present results confirm that the regulation of PLC genes’ expression is crucial in the inflammatory activation of HUVEC. Our results indicate that in HUVEC selected PLC genes, namely PLCB1, PLCB3, PLCB4, PLCD1 and PLCG1 behave oppositely under LPS or Oleuropein stimulation.
In previous reports, PLC β1 was involved in inflammatory stimulation, both in HUVEC (Lo Vasco et al. 2013a, b, c) and in astrocytes (Lo Vasco et al. 2010a), in a complex manner, not completely highlighted. Moreover, PLC β1 was down-regulated in endometriosis compared to normal endometrium (Lo Vasco et al. 2012a, b). PLC β3 was also claimed to be affected with inflammatory stimulation. In fact, PLC β3 underwent down-regulation during astrocyte LPS-activation (Lo Vasco et al., 2010a, b) and up-regulation in endometriosis (Lo Vasco et al. 2012a, 2012b).
Previous studies indicated that PLC γ isoforms are involved in the inflammatory activation of macrophages (Di Raimo et al. 2016). PLC γ subfamily is activated in the presence of AA that serves as a common link between receptor activation of Phospholipase A2 and the PI pathway (Suh et al. 2008). PLC γ isoforms are also involved in the activation of EC by the vascular endothelial growth factor (VEGF) via its receptor VEGFR-2 (Baker et al. 2009). Plc γ1 was suggested to be involved in astrocyte LPS-induced activation (Lo Vasco et al. 2010a). Literature data demonstrated PLC γ1 to be essential for T cell development, activation and tolerance. In fact, in both in vitro and animal models PLC γ1 affects the activities of regulatory T cells, causing inflammatory/autoimmune symptoms (Kalra et al. 2004; Fu et al. 2010).
The involvement of PLC δ1 in inflammatory diseases was also suggested. In fact, PLC δ1 isoform was down-regulated in endometriosis (Lo Vasco et al. 2012a, 2012b) and lack of the Plcd1 gene in knock-out (KO) mice induces skin inflammation (Suh et al. 2008).
Beside confirming the recruitment after inflammatory stimulation, our findings suggest that the expression of PLC genes can be finely regulated by using anti-inflammatory molecules. Our results suggest that Oleuropein can affect the LPS induced modifications of PLC expression. That supports the hypothesis of a possible role as a modulation molecule which can regulate the expression of signalling elements during inflammation. This observation paves the way to novel insights to the regulation and the possible therapeutic use of signalling molecules in the control of the inflammation cascade.
The intracellular localization of PLC enzymes was also modified by different treatments (Figs. 2, 3 and 4). The translocation of PLC enzymes from cytoplasm to nucleus and vice versa was reported and might depend on the cell cycle (Suh et al. 2008). The present results indicate that PLC γ1 and PLC η1 enzymes were located within the cytoplasm of untreated HUVEC and in the nucleus of LPS treated HUVEC (Fig. 4). Although the mRNA for PLCD4 gene was not detected, the corresponding protein PLC δ4 was detected in the nucleus of untreated HUVEC, while it was not detected in LPS treated HUVEC (Fig. 3). PLC γ2 was not detected in both untreated HUVEC and LPS treated HUVEC. Surprisingly, PLC γ2 was detected in the nuclei of HUVEC treated with LPS plus Blueberry extract, as well as in the perinuclear halo of HUVEC cells treated with LPS plus Oleuropein (Fig. 4).
We also observed podosome-like structures in differently LPS treated HUVEC (Fig. 5). Interestingly, in podosomes we detected PLC β1 and PLC η2. Migration and invasion of EC are the trigger point of vessels’ formation and integrity (Osiak et al. 2005). Podosomes are specialized microdomains of the plasma membrane consisting of a complex structure with a central actin core (Rottiers et al. 2009). Exclusively intact EC can form podosomes. During cell migration, podosomes are supposed to confer adhesive and invasive properties to cells. RhoGTPases and Src kinase signalling regulate the cytoskeleton and, probably, the migration/invasion process. Interestingly, PLC β enzymes are regulated by heterotrimeric GTP-binding proteins as well as PLC η enzymes are activated through GPCR stimulation (Suh et al. 2008). Therefore, the presence of both PLC β1 and PLC η2 in HUVEC podosomes, which might indicate the involvement in cytoskeleton remodeling, will require further investigations.
Acknowledgements
The authors wish to thank the “Serena Talarico Association” for encouragement and Dr. Ryan Alejandro Cavanaugh for technical assistance.
References
- Alarcon de la Lastra C, Barranco MD, Motilva V, Herrerıas JM. Mediterranean diet and health: biological importance of olive oil. Curr Pharm Des. 2001;7(10):933–950. doi: 10.2174/1381612013397654. [DOI] [PubMed] [Google Scholar]
- Baker N, O'Meara SJ, Scannell M, Maderna P, Godson C (2009) Lipoxin A4: anti-inflammatory and anti-angiogenic impact on endothelial cells. J Immunol 15;182(6):3819–26. doi: 10.4049/jimmunol.0803175 [DOI] [PubMed]
- Berridge MJ, Dupont G. Spatial and temporal signalling by calcium. Curr Opin Cell Biol. 1994;6(2):267–274. doi: 10.1016/0955-0674(94)90146-5. [DOI] [PubMed] [Google Scholar]
- Bunney TD, Katan M. PLC regulation: emerging pictures for molecular mechanisms. Trends Biochem Sci. 2011;36(2):88–96. doi: 10.1016/j.tibs.2010.08.003. [DOI] [PubMed] [Google Scholar]
- Carmeliet P. Angiogenesis in health and disease. Nat Med. 2003;9:653–660. doi: 10.1038/nm0603-653. [DOI] [PubMed] [Google Scholar]
- Cesa S, Carradori S, Bellagamba G, Locatelli M, Casadei M A, Masci A, Paolicelli P (2017) Evaluation of processing effects on anthocyanin content and colour modifications of Blueberry (Vaccinium spp.) extracts: comparison between HPLC-DAD and CIELAB analyses. Food Chemistry, published online doi: 10.1016/j.foodchem.2017.03.153 [DOI] [PubMed]
- Christ M, Wehling M. Cardiovascular steroid actions: swift swallows or sluggish snails? Cardiovasc Res. 1998;40:34–44. doi: 10.1016/S0008-6363(98)00147-3. [DOI] [PubMed] [Google Scholar]
- Comer FI, Parent CA. Phosphoinositides specify polarity during epithelial organ development. Cell. 2007;128(2):239–240. doi: 10.1016/j.cell.2007.01.010. [DOI] [PubMed] [Google Scholar]
- Del Rio D, Rodriguez-Mateos A, Spencer J, Tognolini M, Borges G, Crozier A. Dietary (poly)phenolics in human health: structures, bioavailability, and evidence of protective effects against chronic diseases. Antioxid Redox Signal. 2013;18:1818–1892. doi: 10.1089/ars.2012.4581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Raimo T, Leopizzi M, Mangino G, Rocca CD, Businaro R, Longo L, Lo Vasco VR. Different expression and subcellular localization of phosphoinositide-specific phospholipase C enzymes in differently polarized macrophages. J Cell Commun Signal. 2016;10(4):283–293. doi: 10.1007/s12079-016-0335-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Divecha N, Irvine RF. Phospholipid signaling. Cell. 1995;80(2):269–278. doi: 10.1016/0092-8674(95)90409-3. [DOI] [PubMed] [Google Scholar]
- Duymuş HG, Göger F, Başer KHC. In vitro antioxidant properties and anthocyanin compositions of elderberry extracts. Food Chem. 2014;155:112–119. doi: 10.1016/j.foodchem.2014.01.028. [DOI] [PubMed] [Google Scholar]
- Elsner J, Dichmann S, Dobos GJ, Kapp A. Actin polymerization in human eosinophils, unlike human neutrophils, depends on intracellular calcium mobilization. J Cell Physiol. 1996;167:548–555. doi: 10.1002/(SICI)1097-4652(199606)167:3<548::AID-JCP18>3.0.CO;2-#. [DOI] [PubMed] [Google Scholar]
- Ferrara N, Davis-Smyth T. The biology of vascular endothelial growth factor. Endocr Rev. 1997;18:4–25. doi: 10.1210/edrv.18.1.0287. [DOI] [PubMed] [Google Scholar]
- Fu G, Chen Y, Yu M, Podd A, Schuman J, He Y, Di L, Yassai M, Haribhai D, North PE, Gorski J, Williams CB, Wang D, Wen R. Phospholipase C{gamma}1 is essential for T cell development, activation, and tolerance. J Exp Med. 2010;207(2):309–318. doi: 10.1084/jem.20090880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garzón GA, Narváez CE, Riedl KM, Schwartz SJ. Chemical composition, anthocyanins, non-anthocyanin phenolics and antioxidant activity of wild bilberry (Vaccinium Meridionale Swartz) from Colombia. Food Chem. 2010;122:980–986. doi: 10.1016/j.foodchem.2010.03.017. [DOI] [Google Scholar]
- Giampieri F, Alvarez-Suarez JM, Mazzoni L, Forbes-Hernandez TY, Gasparrini M, Gonzàlez-Paramàs AM, Santos-Buelga C, Quiles JL, Bompadre S, Mezzettia B, et al. An anthocyanin-rich strawberry extract protects against oxidative stress damage and improves mitochondrial functionality in human dermal fibroblasts exposed to an oxidizing agent. Food Funct. 2014;5:1939–1948. doi: 10.1039/C4FO00048J. [DOI] [PubMed] [Google Scholar]
- Giampieri F, Alvarez-Suarez JM, Mazzoni L, Forbes-Hernandez TY, Gasparrini M, Gonzàlez-Paramàs AM, Santos-Buelga C, Quiles JL, Bompadre S, Mezzettia B, et al. Polyphenol-rich strawberry extract protects human dermal fibroblasts against hydrogen peroxide oxidative damage and improves mitochondrial functionality. Molecules. 2014;19:7798–7816. doi: 10.3390/molecules19067798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gloe T, Pohl U. Laminin binding conveys mechanosensing in endothelial cells. News Physiol Sci. 2002;17:166–169. doi: 10.1152/nips.01381.2001. [DOI] [PubMed] [Google Scholar]
- Grienddling KK, Alexander WR. Endothelial control of the cardiovascular system: recent advances. FASEB J. 1996;10:283–292. doi: 10.1096/fasebj.10.2.8641561. [DOI] [PubMed] [Google Scholar]
- Hisatsune C, Nakamura K, Kuroda Y, Nakamura T, Mikoshiba K. Amplification of Ca2+ signaling by diacylglycerolmediatedinositol 1,4,5-trisphosphate production. J Biol Chem. 2005;280(12):11723–11730. doi: 10.1074/jbc.M409535200. [DOI] [PubMed] [Google Scholar]
- Kalra R, Singh SP, Pena-Philippides JC, Langley RJ, Razani-Boroujerdi S, Sopori ML. Immunosuppressive and anti-inflammatory effects of nicotine administered by patch in an animal model. Clin Diagn Lab Immunol. 2004;11(3):563–568. doi: 10.1128/CDLI.11.3.563-568.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li C, Feng J, Huang W, An X. Composition of polyphenols and antioxidant activity of Rabbiteye Blueberry (Vaccinium ashei) in Nanjing. J Agric Food Chem. 2013;61(3):523–531. doi: 10.1021/jf3046158. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR. Role of phosphoinositide-specific phospholipase C 2 in isolated and syndromic mental retardation. Eur Neurol. 2011;65:264–269. doi: 10.1159/000327307. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR. 1p36.32 rearrengements and the role of PI-PLC 2 in nervous tumours. J Neuro-Oncol. 2011;103(3):409–416. doi: 10.1007/s11060-010-0422-3. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Fabrizi C, Artico M, Cocco L, Billi AM, Fumagalli L, Manzoli FA. Expression of phosphoinositide-specific phospholipase C isoenzymes in cultured astrocytes. J Cell Biochem. 2007;100(4):952–959. doi: 10.1002/jcb.21048. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Fabrizi C, Panetta B, Fumagalli L, Cocco L. Expression pattern and sub cellular distribution of phosphoinositide specific phospholipase C enzymes after treatment with U-73122 in rat astrocytoma cells. J Cell Biochem. 2010;110(4):1005–1012. doi: 10.1002/jcb.22614. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Fabrizi C, Fumagalli L, Cocco L. Expression of phosphoinositide specific phospholipase C isoenzymes in cultured astrocytes activated after stimulation with lipopolysaccharide. J Cell Biochem. 2010;109(5):1006–1012. doi: 10.1002/jcb.22480. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Pacini L, Di Raimo T, D’Arcangelo D, Businaro R. Expression of phosphoinositide-specific phospholipase C isoforms in HUVEC. J Clin Pathol. 2011;64(10):911–915. doi: 10.1136/jclinpath-2011-200096. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Leopizzi M, Chiappetta C, Businaro R, Polonia P, Della Rocca C, Litta P. Expression of phosphoinositide-specific phospholipase C enzymes in normal endometrium and in endometriosis. Fertil Steril. 2012;98(2):410–414. doi: 10.1016/j.fertnstert.2012.04.020. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Cardinale G, Polonia P. Deletion of PLCB1 gene in schizophrenia affected patients. J Cell Mol Med. 2012;16(4):844–851. doi: 10.1111/j.1582-4934.2011.01363.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo Vasco VR, Leopizzi M, Chiappetta C, Puggioni C, Della Rocca C, Businaro R. Lypopolysaccharide down-regulates the expression of selected phospholipase C genes in cultured endothelial cells. Inflammation. 2013;36(4):862–868. doi: 10.1007/s10753-013-9613-3. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Leopizzi M, Chiappetta C, Puggioni C, Della Rocca C, Businaro R. Lypopolysaccharide down-regulates the expression of selected phospholipase C genes in cultured endothelial cells. Inflammation. 2013;36(4):862–868. doi: 10.1007/s10753-013-9613-3. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Longo L, Polonia P. Phosphoinositide-specific phospholipase C ß1 gene deletion in bipolar disorder affected patient. J Cell Commun Signal. 2013;7(1):25–29. doi: 10.1007/s12079-012-0182-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo Vasco VR, Leopizzi M, Puggioni C, Della Rocca C. I, Businaro R. Neuropeptide Y significantly reduces the expression of PLCB2, PLCD1 and moderately decreases selected PLC genes in endothelial cells. Mol Cell Biochem. 2014;394(1–2):43–52. doi: 10.1007/s11010-014-2079-2. [DOI] [PubMed] [Google Scholar]
- Lo Vasco VR, Leopizzi M, Puggioni C, Della Rocca C, Businaro R. Fibroblast growth factor acts upon the transcription of phospholipase C genes in human umbilical vein endothelial cells. Mol Cell Biochem. 2014;388(1):51–59. doi: 10.1007/s11010-013-1898-x. [DOI] [PubMed] [Google Scholar]
- Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods. 1993;65:55–63. doi: 10.1016/0022-1759(83)90303-4. [DOI] [PubMed] [Google Scholar]
- Nakamura Y, Fukami K. Roles of phospholipase C isozymes in organogenesis and embryonic development. Physiology. 2009;24:332–341. doi: 10.1152/physiol.00031.2009. [DOI] [PubMed] [Google Scholar]
- Neto C. Cranberry and Blueberry: evidence for protective effects against cancer and vascular diseases. Mol Nutr Food Res. 2007;51:652–664. doi: 10.1002/mnfr.200600279. [DOI] [PubMed] [Google Scholar]
- Osiak A, Zenner G, Lined S. Subconfluent endothelial cells form podosomes downstream of cytokine and RhoGTPase signalling. Exp Cell Res. 2005;307:342–353. doi: 10.1016/j.yexcr.2005.03.035. [DOI] [PubMed] [Google Scholar]
- Revest PA, Abbott NJ, Gillespie JI. Receptor mediated changes in intracellular [Ca2+] in cultured rat brain capillary endothelial cells. Brain Res. 1991;549:159–161. doi: 10.1016/0006-8993(91)90614-2. [DOI] [PubMed] [Google Scholar]
- Rodrigo R, Libuy M, Feliu F, Hasson D. Polyphenols in disease: from diet to supplements. Curr Pharm Biotechnol. 2014;15:304–317. doi: 10.2174/138920101504140825113815. [DOI] [PubMed] [Google Scholar]
- Rottiers P, Saltel F, Daubon T, Chaigne-Delalande B, Tridon V, Billottet C, Reuzeau E, Génot E (2009) TGFbeta-induced endothelial podosomes mediate basement membrane collagen degradation in arterial vessels. J Cell Sci 1;122(Pt 23):4311–8. doi: 10.1242/jcs.057448 [DOI] [PubMed]
- Seeram NP, Adams LS, Zhang Y, Lee R, Sand D, Scheuller HS, Heber D. Blackberry, black Raspberry, Blueberry, cranberry, red Raspberry, and strawberry extracts inhibit growth and stimulate apoptosis of human cancer cells in vitro. J Agric Food Chem. 2006;54:9329–9339. doi: 10.1021/jf061750g. [DOI] [PubMed] [Google Scholar]
- Slatnar A, Jakopic J, Stampar F, Veberic R, Jamnik P. The effect of bioactive compounds on in vitro and in vivo antioxidant activity of different berry juices. PLoS One. 2012;7:10. doi: 10.1371/journal.pone.0047880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stefani M, Rigacci S. Beneficial properties of natural phenols: highlight on protection against pathological conditions associated with amyloid aggregation. Biofactors. 2014;40:482–493. doi: 10.1002/biof.1171. [DOI] [PubMed] [Google Scholar]
- Suh PG, Park J, Manzoli L, Cocco L, Peak JC, Katan M, Fukami K, Kataoka T, Yuk S, Ryu SH. Multiple roles of phosphoinositide-specific phospholipase C isozymes. BMB Rep. 2008;41:415–434. doi: 10.5483/BMBRep.2008.41.6.415. [DOI] [PubMed] [Google Scholar]
- Wedge DE, Meepagala KM, Magee JB, Hope Smith S, Huang G, Larcom LL. Anticarcinogenic activity of strawberry, Blueberry, and Raspberry extracts to breast and cervical cancer cells. J Med Food. 2001;4:49–51. doi: 10.1089/10966200152053703. [DOI] [PubMed] [Google Scholar]