Fig. 1.
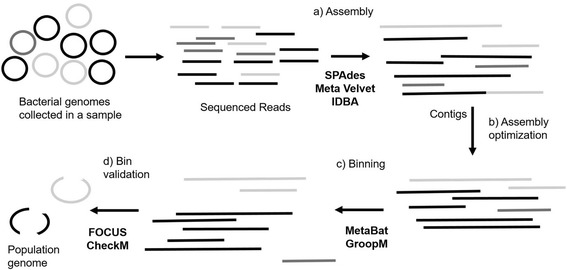
Overview of the workflow developed with the tools applied at each step (in bold). a Metagenomic reads are assembled using three assemblers: SPAdes, MetaVelvet, and IDBA. b Optimization of assembly tool using assembly statistics. c Assembled contigs from optimal assembly were binned using: MetaBat, and GroopM. d Optimal binning tool selected through bin validation. Colors (black, dark grey, and light grey) depict different microbial species, each line of a color representing the sequence belonging to a bacterial species
