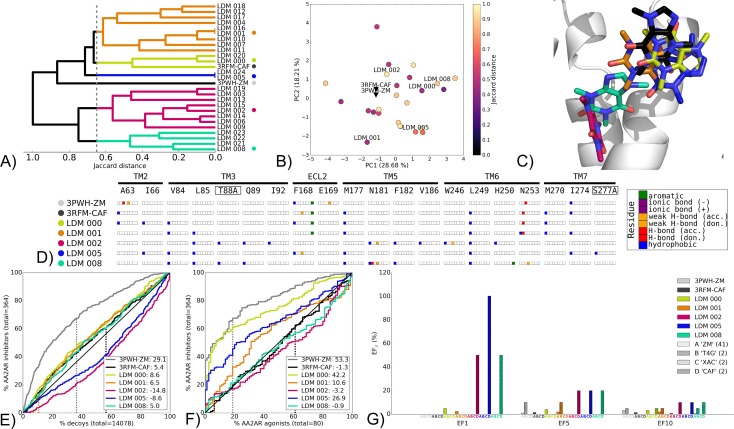
Fig 6. Chemotype switch LDM experiment on the AA2AR from 3PWH-ZM to 3RFM-CAF, using CAF as a refinement ligand.
A) Dendrogram of the top 25 LDM models and X-ray structures, a cutoff line identifies different LDM clusters and their representative LDM models are designated by a colored dot. Representative LDM models are the highest scoring within the cluster based on the OPUS-ICM metric. B) Comparison of binding pocket conformation between the top 25 LDM models and X-ray structures. LDM models are colored based on their IFP Jaccard distance with the destination X-ray structure, 3RFM-CAF. C) Binding poses of the representative LDM models and the destination X-ray structure with the X-ray ligand in black and the LDM models coloured based on their representative clusters defined in A. D) IFP of the representative LDM models and the X-ray structures. Interaction type is described for each residue of the binding pocket: hydrophobic interaction, hydrogen bond (H-bond) donor and acceptor, weak hydrogen bond (weak H-bond) donor and acceptor, ionic bond positive (+) and negative (-) and aromatic interaction. VS performance is described with ROC curves to visualise E) the recovery of AA2AR inhibitors vs. decoys and F) the selectivity of AA2AR inhibitors over AA2AR agonists. X-ray structures are in black, with the LDM models coloured based on their relative clusters identified in A. The relative rank of the LDM refinement ligand is identified with a vertical dashed line. This vertical line for some models may be masked by others if the ligand is very highly ranked. The ROC curve figure inset shows NSQ_AUC values for each binding pocket. G) bar chart is used to visualise the EF for representative AA2AR inhibitor chemotypes at EF1, EF5 and EF10. Chemotypes A ‘ZM-like’, B ‘T4G-like’, C ‘XAC-like’ and D ‘CAF-like’ represent only a subset of AA2AR inhibitors ligands (S2B Fig). The EF bar chart inset shows the number of ligands for each chemotype cluster between parenthesis. Origin and destination X-ray structure chemotype EF shown in grey and black bars, respectively, with the LDM models coloured based on their relative clusters identified in A.