Fig. 7.
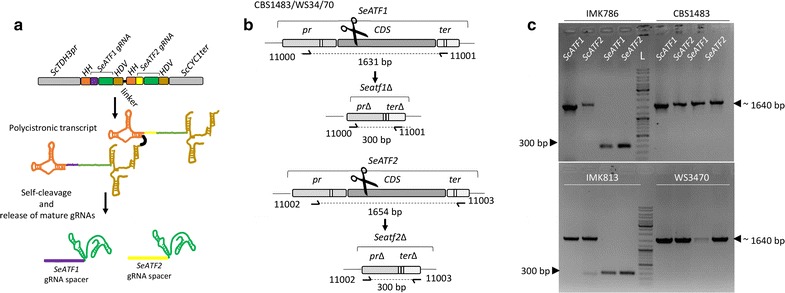
Simultaneous deletion of all SeATF1 and SeATF2 alleles using a single ribozymes flanked gRNA array in S. pastorianus CBS1483 and Weihenstephan 3470 (WS3470). a Representation of the gRNA array expression cassette in pUDP044. The dual gRNA array was under the control of the RNA polymerase II promoter ScTDH3 and ScCYC1 terminator. Each gRNA was flanked on its 5′ by a hammerhead ribozyme (HH represented in orange) and on its 3′ by a hepatitis delta virus (HDV represented in bronze) ribozyme and they were separated by a 10 bp linker. Upon ribozyme self-cleavage, the mature gRNAs are released. The SeATF1 guiding spacer (in purple), the SeATF2 guiding spacer (in yellow) and the constant structural gRNA fragment (in green) are indicated. b Schematic representation of the SeATF1 and SeATF2 editing upon transformation of CBS1483 with pUDP044. The primers for the validation of transformants are indicated. c Validation of transformants of the S. pastorianus CBS1483 strain with pUDP044 in presence of a 120 bp repair DNA. The PCR reactions were performed with the primers pairs 11000/11001 for SeATF1 and 11002/11003 for SeATF2. The isolate renamed IMK786 exhibited bands at 300 bp corresponding to the deletions of SeATF1 and SeATF2. ScATF1 and ScATF2 were amplified using the primer pairs 10996/10997 and 10998/10999 respectively and exhibited wild type length. Similarly, transformants resulting from the transformation of pUDP044 in presence of a 120 bp repair DNA were checked with the primers pairs 11000/11001 for SeATF1 and 11002/11003 for SeATF2. The isolate renamed IMK813 exhibited bands at 300 bp corresponding to the deletions of SeATF1 and SeATF2
