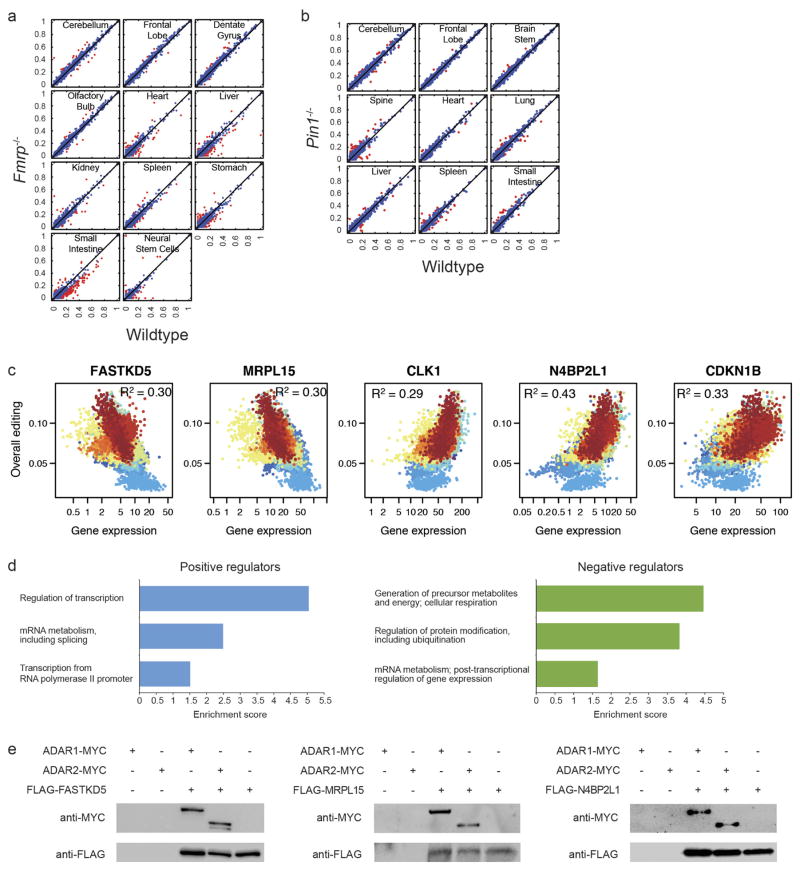
Extended Data Figure 9. Analysis of FMRP, PIN1 and other potential regulators of RNA editing.
a, Comparison of average editing levels in 10 tissues and neural stem cells of wild-type and Fmrp−/− mice at reproducible sites (s.d. <10% in wild-type and Fmrp−/− replicates). Sites that differ by more than 10% in editing levels between wild-type and Fmrp−/− mice are marked in red. b, Comparison of average editing levels in 9 tissues of wild-type and Pin1−/− mice at reproducible sites (s.d. <10% in wild-type and Pin1−/− replicates). Sites that differ by more than 10% in editing levels between wild-type and Pin1−/− mice are marked in red. c, Correlation of the expression levels of the top negative (FASTKD5 and MRPL15) or positive (CLK1, N4BP2L1 and CDKN1B) candidate regulators with overall editing of all sites in the GTEx samples. R2 values were calculated by robust linear regressions on overall editing levels and logarithmic transformed RPKM values. d, GO analysis of the 144 putative positive regulators and 147 putative negative regulators of editing. The top three biological processes that are reported by both DAVID and Panther are given for each set of regulators. e, Both ADAR1 and ADAR2 co-immunoprecipitates with FASTKD5, MRPL15 and N4BP2L1. HEK293T cell lysates were incubated with anti-Flag M2 beads to immunoprecipitate each regulator and concurrently pull down the ADAR enzymes.