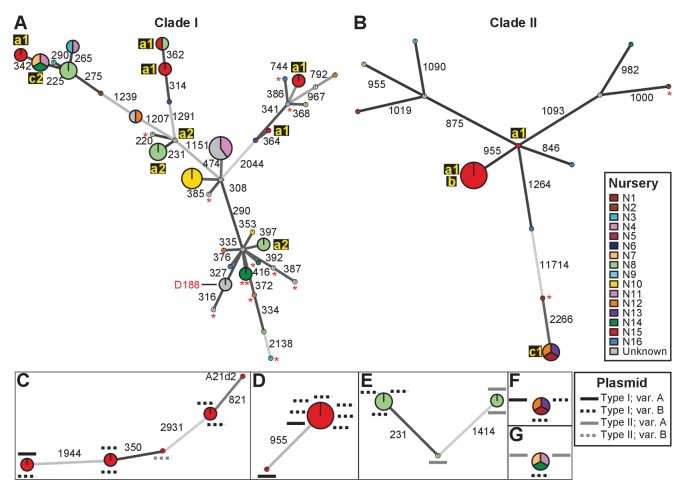
Figure 2. Analysis of SNPs reveals three transmission patterns of pathogenic Rhodococcus.
Minimum spanning networks of isolates of (A) Clade I and (B) Clade II. Each genotype is displayed as a circle, with sizes scaled to represent the number of associated isolates (smallest = 1 isolate). Colors represent the source of the isolates (see key), with coloring proportional to the ratio of isolates from each source. Lower-case letters and numbers (a1, a2, b, c1, and c2) highlight potential transmission patterns; see panels C–G). Asterisks = lacking virulence genes. The genotype that includes D188 is indicated. (C, D) Minimum spanning networks of pathogenic isolates belonging to Clade I (C; ‘a1’) and Clade II (D; ‘a1’ and ‘b’) from nursery N15. A21d2 lacks a virulence plasmid and its virulence loci are present in the chromosome. (E) The minimum spanning network for isolates of pathogenic isolates from nursery N8 (‘a2’). (F) The epidemiological link ‘c1’ between isolates from nurseries N1, N12, and N13. (G) The epidemiological link ‘c2’ between isolates from nurseries N7, N11, and N14. Plasmid types and their variants are mapped onto each of the nodes (see key). Numbers adjacent to connecting lines indicate the number of SNPs that separate each genotype. The lengths of connecting lines are arbitrary; gray lines indicate distances that exceed an arbitrary threshold.