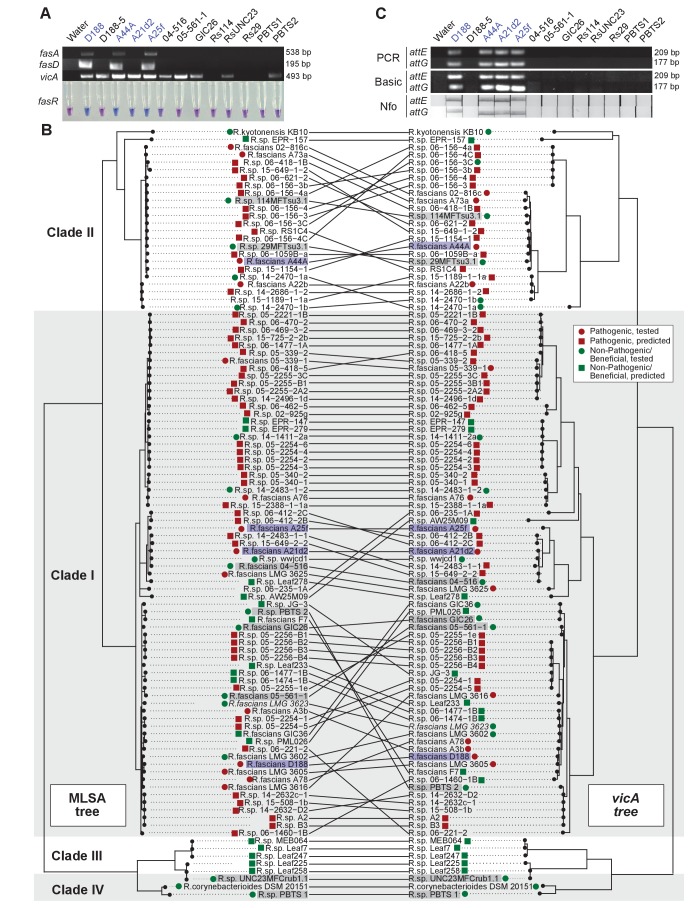
Figure 8. Use of the vicA gene does not discriminate pathogenic Rhodococcus.
(A) (Top) Using locus-specific primers and DNA from the listed isolates, fragments of fasA, fasD and vicA were PCR amplified. Pathogenic isolates are labeled in blue; virulence-gene-lacking isolates are labeled in black. Products were resolved on a 1% TAE agarose gel. Amplicon sizes are listed to the right of the gel images. (Bottom) LAMP assay detection of fasR from the same DNA samples. A positive result is visualized by a blue color. Negative results are light purple. (B) Congruency between species (left) and vicA trees (right). Clades other than I–IV are not shown. Highlighted isolates are the same as in A, pathogenic isolates are highlighted in blue, virulence-gene-lacking isolates are highlighted in gray. (C) Standard endpoint PCR, RPA Basic, and RPA nfo were used to detect attE or attG from DNA extracted from isolates of Rhodococcus. Pathogenic isolates are labeled in blue. For PCR and RPA basic, product sizes are list to the right of the figure. For RPA nfo, the presence of the test band is indicative of a positive reaction; the control bands for all strips were confirmed (not shown).