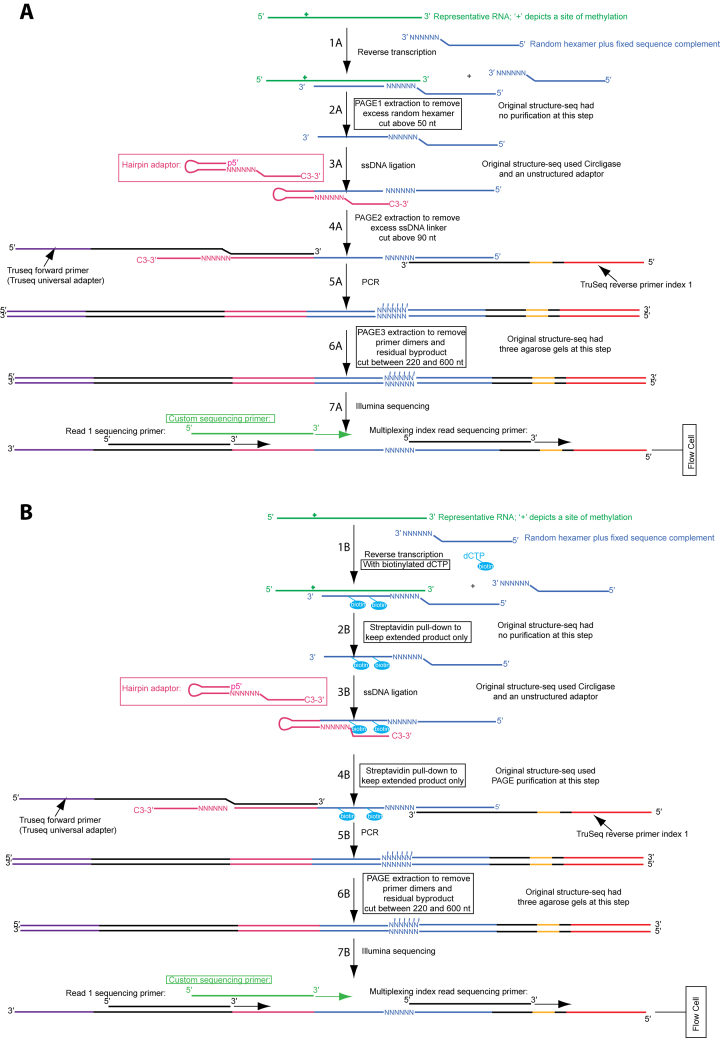
Figure 1.
Two versions of Structure-seq2 produce high quality data. In Structure-seq2, RNA (kelly green) is first modified by DMS or another chemical that can be read-out through reverse transcription. The RNA is then prepared for Illumina NGS sequencing by conversion to cDNA (Step 1A/1B, blue), ligating an adaptor (Step 3A/3B), and amplifying the products while incorporating TruSeq primer sequences (Step 5A/5B). In order to increase library quality, numerous improvements were made to the original Structure-seq protocol (boxed). These include performing the ligation with a hairpin adaptor and T4 DNA ligase (Step 3A/3B; pink) (10), and adding various purification steps to remove a deleterious by-product (Figure 2A). We present two options for purification: PAGE purification (A) or a biotin–streptavidin pull down (B). In the PAGE purification method, an additional PAGE purification step is added after reverse transcription (Step 2A). In the biotin–streptavidin pull down method, biotinylated dNTPs (cyan) are incorporated into the extended product during reverse transcription (Step 1B) and are purified via a magnetic streptavidin pull down after reverse transcription (Step 2B) and after ligation (Step 4B). There is also a common, final PAGE purification step following amplification (Step 5A/5B). Finally, a custom sequencing primer (light green) is used during sequencing (Step 7A/7B) to further provide high quality data. Supplementary Figure S1 is a version of this figure with all the nucleotides shown explicitly.