Fig. 4.
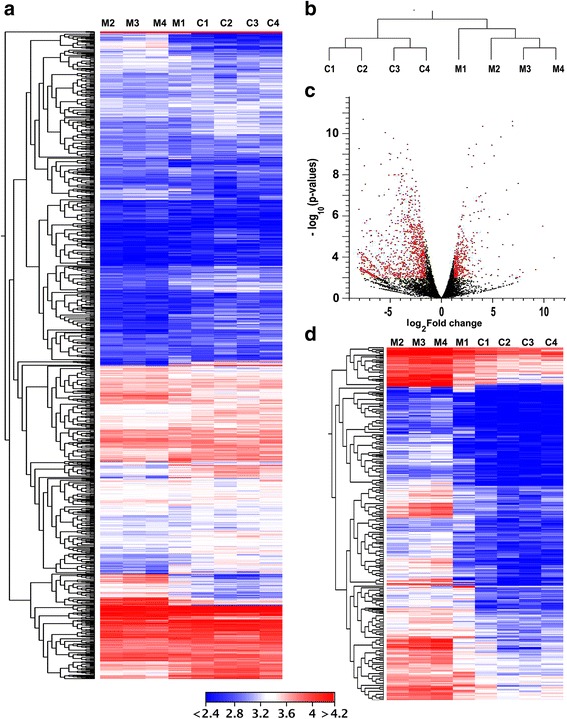
Gene expression analysis of RNA isolated from dsmalE (control, M1-M4) and dsCBP injected (C1-C4) last instar Tribolium larvae using the empirical analysis of DGE (EDGE) algorithm in CLC Genomics Workbench (Version: 9.5.9) and WEGO. a Heatmap showing the overall gene expression pattern of all four biological replicates from malE or CBP dsRNA injected larvae. The color spectrum, stretching from blue to red, represents TMM normalized expression values obtained after EDGE analysis. b Hierarchical clustering based on the overall gene expression pattern showing the relative differences within and among the biological replicates from malE or CBP dsRNA injected larvae. c Volcano plot of expression data after EDGE analysis. In the plot, log2 of fold change between the malE and CBP dsRNA-treated insects are plotted against −log10 (p), where p is a probability value (for a given gene) that is associated with the EDGE comparison of the two groups of samples. The red dots indicate the number of significantly up- and down-regulated genes using p < 0.01 and ±2-fold change as the cutoff threshold. d Heatmap illustrating the TMM normalized expression values of 1306 significantly downregulated genes (p < 0.01, ≤2-fold ↓) after CBP knockdown in Tribolium larvae. Color-coding is the same as in figure (a)
