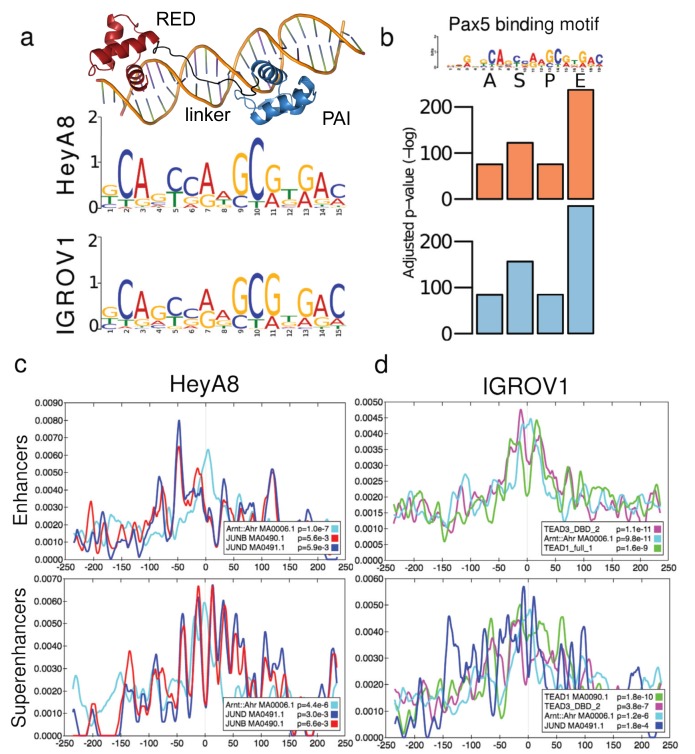
Figure 5. Defining the PAX8 binding motif and identifying candidate co-regulators.
a. A model of the DNA binding domain of PAX8. Binding motifs identified by MEME-ChIP using the H3K27ac positive PAX8 binding sites in the enhancer set for HeyA8 and IGROV1. b. Pax5 binding motif (from Jaspar [43]) was selected as the closest binding motif to the primary PAX8 binding motif based on the adjusted p-value across four sets of PAX8 binding sites used for motif discovery (all (A), superenhancers (S), promoters (P), and enhancer (E) PAX8 peaks) for HeyA8 (orange) and IGROV1 (blue). Significantly enriched motifs within ±250 bp of the summits of PAX8 peaks in c. HeyA8 and (d) IGROV1 cells. Other than PAX-like motifs, only motifs identified in both cell lines are shown (with the exception of TEAD1/3, which was unique to IGROV1). The grey line at 0 bp indicates the summit of the PAX8 peak.