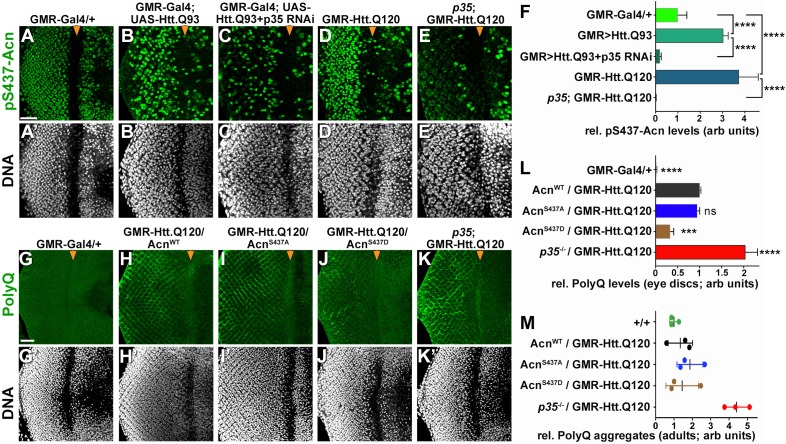
Figure 8. Protein-aggregate-induced phosphorylation of Acn S437 depends on Cdk5/p35.
Projections of confocal micrographs of eye discs stained for pS437-Acn (A–E) or polyQ proteins (G–K) and DNA. Compared to a GMR-Gal4 control (A), GMR-Gal4-driven UAS-Htt.Q93 (B) or GMR-Htt.Q120 (D) expression induced elevated S437 phosphorylation, which was suppressed in eye discs with p35 knockdown (C) or mutant for p35 (E). GMR-directed expression initiates at the furrow (arrowhead) and is more developed toward the posterior. (F) Quantification of S437 phosphorylation (F) averaged constant areas containing about 50 ommatidial clusters located at least 6–8 rows posterior to the furrow. Bar graphs show mean ±SD of integrated densities. Values were normalized to GMR-Gal4 controls and were from one representative experiment out of three repeats. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; for indicated comparisons. Early polyQ accumulation was compared between GMR-Gal4 eye discs as controls (G), and eye discs expressing GMR-Htt.Q120 and carrying a copy of the indicated genomic Acn transgene (H–J) or the p35 null allele (K). Compared to AcnWT (H) and AcnS437A (I) eye discs, in AcnS437D (J) polyQ accumulation was reduced until some 7 to 8 rows of ommatidia posterior to the furrow (arrowhead) and enhanced in p35 mutants (H). Scale bar in A: 40 µm in A-N. Posterior is to the left. (L) Quantification of PolyQ accumulation averaged constant areas containing about 100 ommatidial clusters located at least 2–3 rows posterior to the furrow. Bar graphs show mean ± SD of integrated densities. Values were normalized to AcnWT/GMR-Htt.Q120 (L) and were from one representative experiment out of three repeats. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; compared to AcnWT/GMR-Htt.Q120. (M) Quantification of dot blots measuring aggregated polyQ protein in fly heads expressing GMR-Htt.Q120 and the indicated genomic Acn transgenes or in a p35 mutant background. Control OreR flies (+/+) indicate background level of dot blot measurements (n = 3 biological repeats). The scatter plot shows mean and standard deviation from three separate experiments. Detailed genotypes are listed in Supplementary file 3.