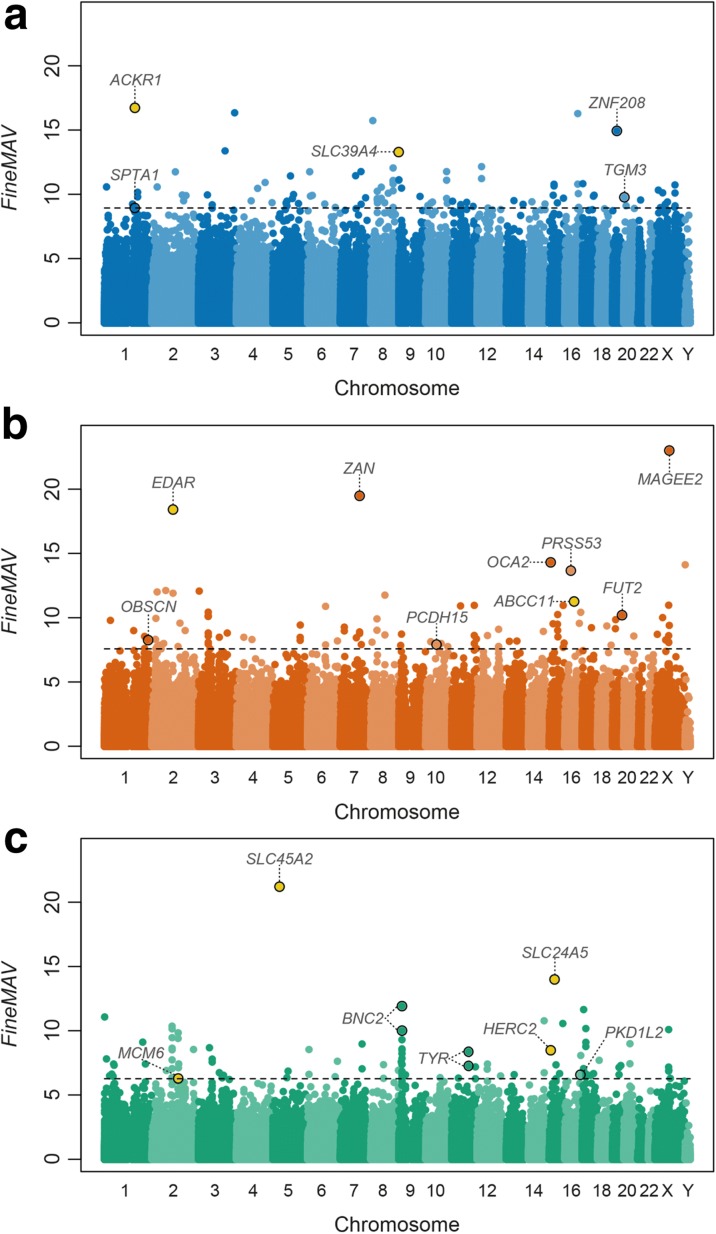
Fig. 4.
Manhattan plot of genome-wide FineMAV scores. FineMAV scores were calculated for genome-wide SNPs from 1000 Genomes Project Phase 3 [37] in three continental populations: (a) Africans (AFR, blue); (b) East Asians (EAS, orange); (c) Europeans (EUR, green). Each dot in the Manhattan plots represents a single SNP plotted according to coordinates in GRCh37. The threshold (dashed lines) was set to include the top 100 variants (top ~ 0.0004% of the whole-genome distribution). All gold-standard SNPs (yellow dots found among the top outliers) and other interesting candidate variants are labeled with the name of the gene they fall into