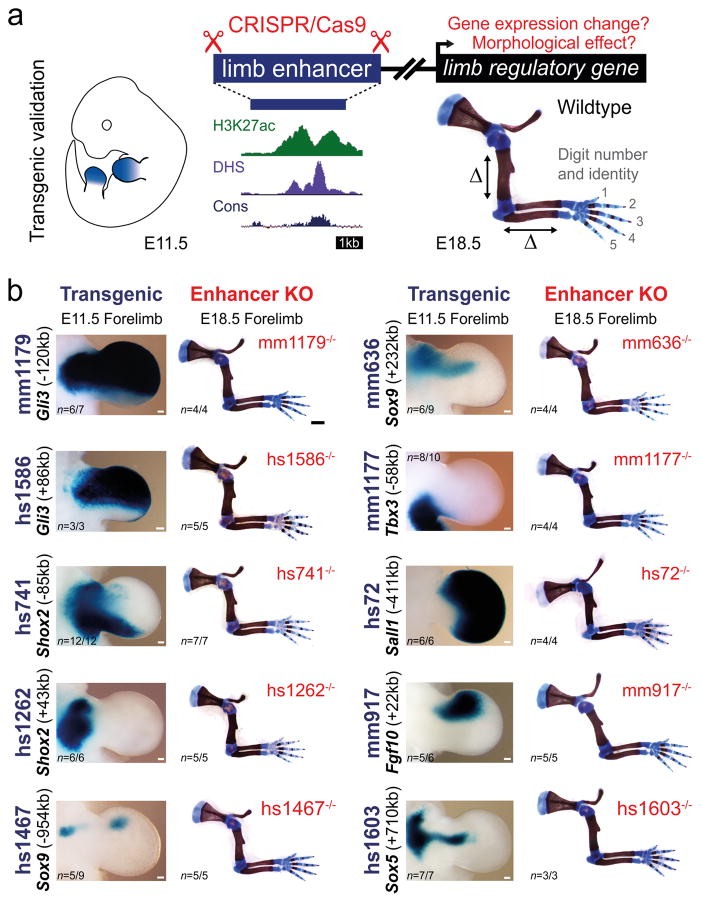
Figure 1. Lack of limb morphological abnormalities in ten enhancer deletion lines.
(a) All selected enhancers are active in the limb mesenchyme (blue shading) at E11.5, are marked by epigenomic H3K27 acetylation and DNase I hypersensitivity (DHS) at E11.5, and contain a conserved core sequence (Cons). Following deletion of individual enhancers (Extended Data Fig. 1a–j), target gene expression and limb morphology were assessed. (b) None on the individual enhancer deletions caused obvious defects in the structure of skeletal elements. Enhancer activities (left, E11.5) and forelimb skeletons of enhancer knockout (KO) embryos (right, E18.5) are shown (see Extended Data Fig. 3 for wild-type controls). Predicted target gene and enhancer distance (+: downstream; -: upstream) from the transcriptional start site (TSS) are indicated. “n”, independent biological replicates with similar results. Scale bars, 100 μm (white), 1 mm (black).