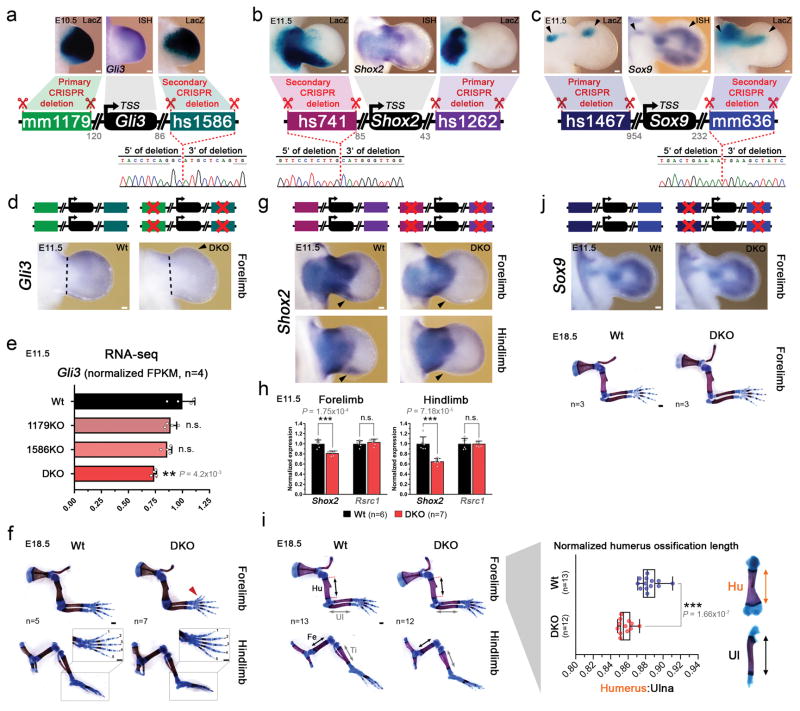
Extended Data Figure 5. Transcriptional and phenotypic impact of dual enhancer deletions engineered by iterative CRISPR/Cas9 genome editing.
(a–c) Upper panels: Enhancer pairs with overlapping limb activities (LacZ), coinciding with domains of predicted target gene expression visualized by in situ hybridization (ISH). For Sox9 enhancers, black arrowheads indicate overlapping domains. Schematics: Double enhancer deletion strategy to delete the three enhancer pairs with overlapping activity (see Methods). Gray numbers indicate enhancer distance (in kb) from the transcriptional start site (TSS). Lower panels: Sanger sequencing verification of the secondary enhancer deletion. Deletion breakpoint is marked by the dashed line. Gray horizontal bars indicate bases present in the primary deletions (single enhancer KO lines, see Extended Data Fig. 1a–j). Shox2- and Sox9- associated LacZ panels are also used in Extended Data Fig. 2. (d) Gli3 transcript distribution (ISH) in wildtype (Wt) and mm1179/hs1586 double enhancer knockout (DKO) embryos. Arrowhead points to reduced Gli3 transcript in the anterior limb mesenchyme. Dashed line indicates dissected hand plate for RNA-seq. (e) RNA-seq confirmed significantly reduced Gli3 expression in hand plates of DKO, but not individual enhancer KO embryos (compared to wildtype hand plates). (f) Unaffected hindlimb morphology in mm1179/hs1586 DKO embryos. Red arrowhead points to digit 1 duplication in forelimbs (see also Fig. 2). (g) Shox2 expression (ISH) in fore- and hindlimbs of hs741/hs1262 double enhancer knockout (DKO) embryos. The distal-posterior domain (arrowhead) is dependent on hs741 (Extended Data Fig. 2). (h) Reduced Shox2 expression in fore- and hindlimbs of hs741/hs1262 DKO embryos (qPCR). Expression of the nearby Rsrc1 gene was unchanged. (i) Left: Representative limb skeletons of wildtype and hs741/hs1262 DKO embryos. Hu, humerus; Ul, ulna; Fe, femur; Ti, Tibia. Right: Mild but significant reduction in humerus ossification length (double arrows) in hs741/hs1262 DKO limb skeletons. ***, P=1.66×10−7 (two-tailed, unpaired t-test). (j) Absence of evident Sox9 expression differences or skeletal abnormalities in embryos lacking both the hs1467 and mm636 enhancers near Sox9. For ISH, transcript distribution was reproduced in at least n=3 independent biological replicates. “n”, number of independent biological replicates with similar results. For bar graphs and boxplots, individual biological replicates are shown as data points. Bar graphs illustrate mean and standard deviation (error bars). Box plot indicates median, interquartile values and range. ***, P < 0.001; **, P < 0.01 (two-tailed, unpaired t-test). n.s., not significant. Scale bars, 100 μm (white) and 500 μm (black).