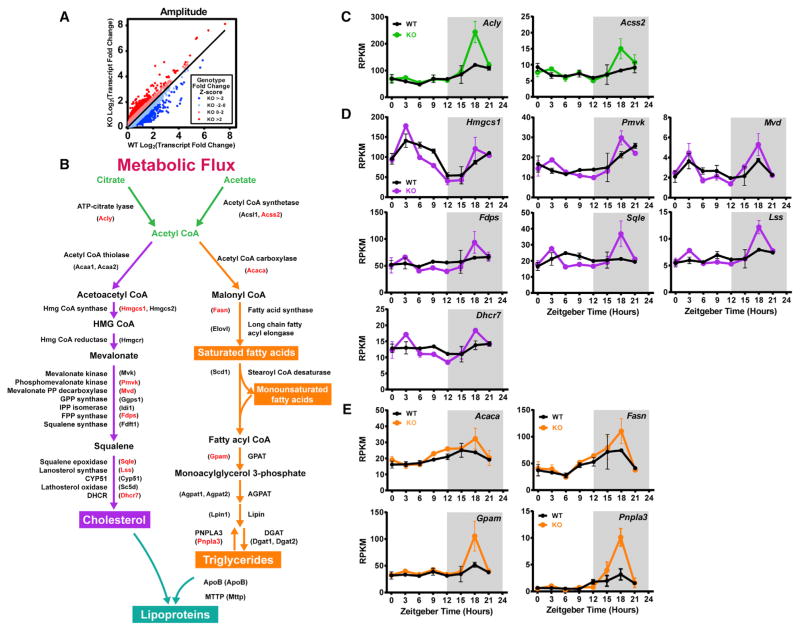
Figure 3. mRNAs Encoding Cholesterol and Lipid Synthesis Enzymes Display Increased Amplitude at ZT18 in the Noct−/− Mouse Liver.
(A) Transcript fold change was determined for each transcript by dividing the maximum expression value by the minimum expression value across time points. Genotype fold change for each transcript was then calculated by dividing the KO transcript fold change by the WT transcript fold change. A Z score was calculated for the genotype fold change and color-coded to highlight the transcripts with the highest amplitude increases in the KO (red) and the lowest amplitude decreases in the KO (blue).
(B) Pathway diagram depicting mRNAs and enzymes involved in the production of acetyl-CoA (green), cholesterol (purple), triglyceride (orange), and lipoproteins (turquoise). The names highlighted with red text had a Z score > 2, as identified in Figures 1I and 1J.
(C–E) mRNA-seq expression plots from WT and Noct−/− (KO) mice for select genes from the metabolic flux pathway in (B) are shown across the circadian cycle (n = 2/genotype/time point). Select genes are color-coded according to their terminal production of Acetyl CoA (green, C), cholesterol (purple, D), or triglycerides (orange, E).
Data points represent the mean ± range.