Fig. 1.
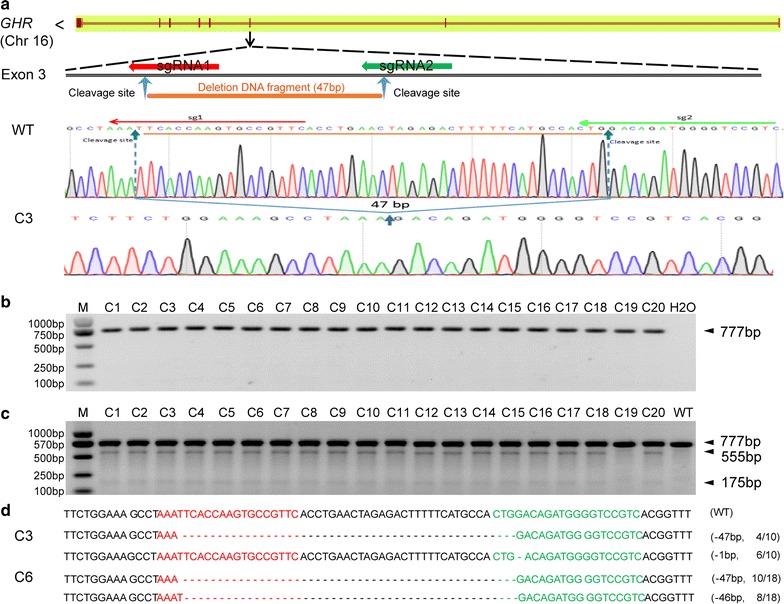
Establishment of GHR-modified single-cell colonies. a Schematic diagram of DNA deletion mediated by the dual-sgRNAs/Cas9 system. The brown line indicates the GHR gene structure from the Ensmbl database. “<” indicates the direction of transcription. The red and green arrows indicate the targeting sites of the designed sgRNAs. The blue arrows indicate cleavage sites mediated by the sgRNAs/Cas9 system. The orange segment and line indicate the deleted 47-bp DNA fragment mediated by the dual-sgRNAs/Cas9 system. WT: wild-type allele. C3: single-cell colony #3. b PCR products harboring the targeted GHR region amplified from pig single-cell colonies (M, DNA maker DL2000; C, colony). c Detection of sgRNAs/Cas9-mediated on-target GHR cleavage by the T7ENI cleavage assay. The band size of wild-type PCR product is 777 bp, and the sizes of T7ENI cut bands are about 555 and 175 bp. The mismatched about 47 bp band was not detected. d Sequence of modified GHR in donor cells for SCNT. The red font indicates the sgRNA1 targeting site, and the green font indicates the sgRNA2 targeting site. C6: single-cell colony #6. “–”: deletion; N/N: positive sequencing out of total sequencing
